Humans Chromosome 1 Fractal Periods Signature is Highly Correlated with Intelligence and Brain Evolution
Abstract
DUF1220 proteins regions show the largest Homo-Sapiens lineage-specific increase in copy number of any protein-coding region in the human genome and map principally to 1q21.1. DUF1220 deletions have been associated with microcephaly and macrocephaly, respectively. DUF1220 copy number has been linked to both brain size in humans and brain evolution among primates. Remarkably, dosage variations involving DUF1220 sequences have now been linked to human brain expansion, autism severity, total IQ, and cognitive and mathematical aptitude scores. We analyzed in chromosome 1q a total of 245 DUF1220 proteins. Finally the method is extended analysing the long 1q21 region from 7 other close primates like Neanderthal, great apes : chimp, gorilla, orangutan and monkeys : macaque, marmoset, vervet. This remarkable property is confirmed by comparing these primates to other mammals such as mice, rabbit, cow, dolphin and Elephant. We then show four classes of multi-periodic fractal structures for all 19 DUF1220 regions and 19 NBPF genes studied cases. The analysis of these spectra of fractal periods1 reveals a simple linear interdependence, hierarchization and unification between the numerical sequences of each of these 4 spectra and the sequences of Fibonacci and Lucas. Given the evidence of this numerical relationship, we suggest that this discovery may be one of the major causes of a cognitive development of man superior to that of the great primates. Finally the mathematical roots of this whole numbers resonance patterns is discussed.
Author Contributions
Academic Editor: Rui-Lin Liu, Xian Jiaotong University, China
Checked for plagiarism: Yes
Review by: Single-blind
Copyright © 2017 Jean-claude PEREZ, et al.
 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Competing interests
The authors have declared that no competing interests exist.
Citation:
Introduction
The exact function of the DUF1220 protein is not known. DUF1220 proteins regions show the largest Homo-Sapiens lineage-specific increase in copy number of any protein-coding region in the human genome and map principally to 1q21.1, and partially also in 1p. DUF1220 deletions have been associated with microcephaly and macrocephaly, respectively. In Colorado University Dr Sikela team established that human genome sequences encoding DUF1220, show a dramatically elevated copy number in the human lineage and variation in DUF1220 copy number has been linked to both brain size in humans and brain evolution among primates 1, 2. Attempts to link genetic and I.Q have been proposed from the 1970s 3. Now, dosage variations involving DUF1220 sequences have now been linked to human brain expansion, autism severity, total IQ, and cognitive and mathematical aptitude scores 4.
There are many more copies of DUF1220 encoded in the human genome compared to the genome of any other species (Table 1). Humans have approximately 270 haploid copies, far more than great apes (90–125 copies), monkeys (25–40 copies), and especially prosimians and non-primate mammals (1–9 copies)
Dr Sikela Lab. demonstrates the hypothesis that increasing copy number of sequences encoding DUF1220 protein domains is a major contributor to the evolutionary increase in brain size, neuron number, and cognitive capacity that is associated with the primate order. They propose that this relationship is restricted to the anthropoid sub-order of primates 5, 6, 7, 8.
Table 1.The distribution of DUF1220 and NBPF gene populations in different mammalian species (6)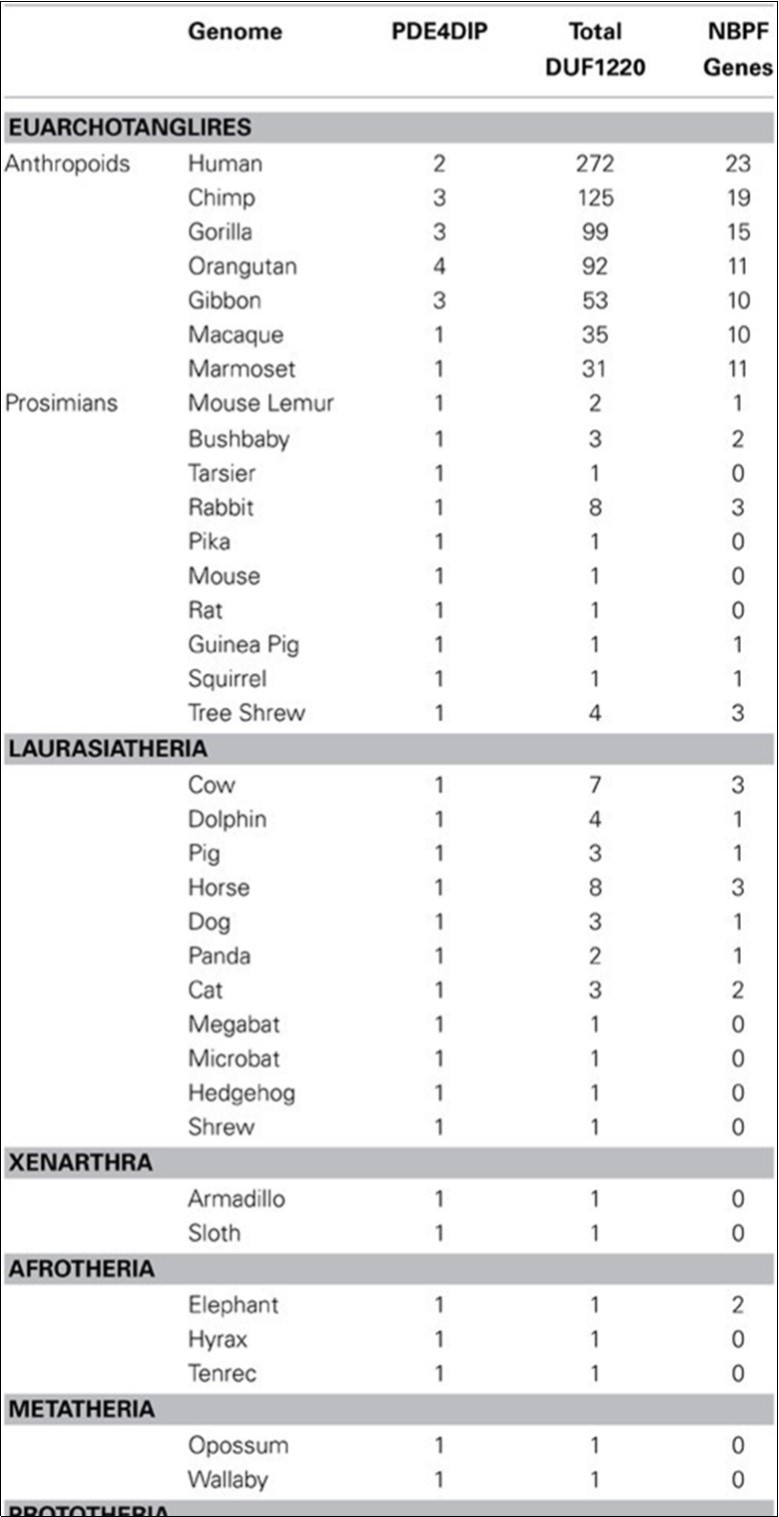
| Dim x Dim y | 0 | d1 1 | d2 | 2 | 3 | …/... d100 4 | 5 | |
| 5 | 318499 | 377103 | 401874 | 416629 | 424005 | |||
| 6 | 369260 | 440557 | 364717 | 405943 | 357653 | |||
| 7 | 389780 | 351783 | 332589 | 318646 | 310074 | |||
| 8 | 253290 | 289225 | 304361 | 258865 | 273966 | |||
| 9 | 275885 | 243343 | 226178 | 259486 | 245390 | |||
| 10 | 188558 | 208306 | 214670 | 218930 | 222246 | |||
| 11 | 208279 | 180867 | 204098 | 187607 | 203513 | |||
| 12 | 220206 | 203013 | 194628 | 190435 | 187589 | |||
| …/... 377 |
| NBPF genes | Resonance | Periods | Fibonacci/Lucas Periods | Shifted sequences | Hyper-reson ance class | ||||||||||||
| Test1 | NBPF1 | 7 | 9 | 16 | 25 | 41 | 5 | 8 | 13 | 21 | 34 | 2 | 1 | 3 | 4 | 7 | B |
| Test2 | NBPF14 | 7 | 9 | 16 | 25 | 41 | 5 | 8 | 13 | 21 | 34 | 2 | 1 | 3 | 4 | 7 | B |
| Test3 | NBPF12 | 7 | 9 | 16 | 25 | 41 | 5 | 8 | 13 | 21 | 34 | 2 | 1 | 3 | 4 | 7 | B |
| Test4NBPF20 | 7 9 16 25 41 | 5 8 13 21 34 | 2 1 3 4 7 | B | |||||||||||||
| Test5NBPF4 | 9 11 20 31 51 | 8 8 16 24 40 = 8x(1 1 2 3 5) | 1 3 47 11 | C | |||||||||||||
| Neanderthal NBPF4 | 9 11 20 31 51 | 8 8 16 24 40 = 8x(1 1 2 3 5) | 1 3 47 11 | C | |||||||||||||
| Test6NBPF10 | 7 9 16 25 41 | 5 8 13 21 34 | 2 1 3 4 7 | B | |||||||||||||
| Test7NBPF6 | 9 11 20 31 51 | 8 8 16 24 40 = 8x(1 1 2 3 5) | 1 3 47 11 | C | |||||||||||||
| Test8NBPF9 | 9 11 20 31 51 | 8 8 16 24 40 = 8x(1 1 2 3 5) | 1 3 47 11 | C | |||||||||||||
| Neanderthal NBPF9 | 9 11 20 31 51 | 8 8 16 24 40 = 8x(1 1 2 3 5) | 1 3 47 11 | C | |||||||||||||
| Test9NBPF9 | 7 9 16 25 41 | 5 8 13 21 34 | 2 1 3 4 7 | B | |||||||||||||
| Test10NBPF9 | 7 9 16 25 41 | 5 8 13 21 34 | 2 1 3 4 7 | B | |||||||||||||
| Test11NBPF6 | 9 11 20 31 51 | 8 8 16 24 40 = 8x(1 1 2 3 5) | 1 3 47 11 | C | |||||||||||||
| Test12NBPF15 | 5 7 12 19 31 | 8 13 21 34 | 1 1 2 3 | A | |||||||||||||
| Neanderthal NBPF15 | 5 7 12 19 31 | 8 13 21 34 | 1 1 2 3 | A | |||||||||||||
| Test13NBPF15 | 5 7 12 19 31 | 8 13 21 34 | 1 1 2 3 | A | |||||||||||||
| Test14NBPF3 | 7 9 16 25 41 | 5 8 13 21 34 | 2 1 3 4 7 | B | |||||||||||||
| Test15NBPF3 | 7 9 16 25 41 | 5 8 13 21 34 | 2 1 3 4 7 | B | |||||||||||||
| Test16NBPF3 | 7 9 16 25 41 | 5 8 13 21 34 | 2 1 3 4 7 | B | |||||||||||||
Indeed, thanks to the CRISPR technology, it is now possible to modify locally the genomes, and more particularly the human genome 9.
Especially since it has been established that this region is extremely fragile, difficult to sequence, and often causes severe cerebral disorders.
On the other hand, the fractal and global structures of the human genome were demonstrated 10.
For more than 25 years, we have been looking for possible global, even digital, structures that would organize DNA, genes, chromosomes, and even whole genomes 11, 12, 13, 14.
We will analyze here several major sequences containing these proteins DUF1220 according to an original approach highlighting kinds of "fractal periodicities". 15, this method shows evidence of "fractal periods" and "resonance periods" characterizing each of the 24 human chromosomes as well as any partial or complete sequence of any chromosome. For example, as illustrated in Figure 1 below, these resonances make it possible to differentiate the respective genomes of Neanderthal and Sapiens on the global scale of the 1q21.1 DUF1220 main rich region.
Figure 1.The 2 respective Neanderthal and Sapiens 1q21.1 DUF1220 region6 have a "resonance" of 12 bp, however, these two radically different resonance curves illustrate a major differentiation of the 2 human species on the GLOBAL scale.
Methods
Analysed Whole Human Genomes :
We have completely and systematically analyzed the DUF1220 protein rich regions in many genomes and more specifically in each of the following 2 reference genomes:
Neanderthal reference genome :-Neanderthalgenome(2014)ref 16
Sapiens HG38 (2013) human reference genome ref 17
- Sapiens HG38 (2013) genomehttps:/ www.ncbi.nlm.nih.gov/grc/human
Analysed DUF1220 regions in Sapiens HG38 chromosome1 :
For more details, please visit complementary materials (part I).
Computing Fractal Periods and Resonances Summary
We introduce here a method of global analysis of the roughness or fractal texture of the DNA sequences at the chromosome scale. To do this, we generalize the method of numerical analysis of the "Master Code of Biology" 15. Thus, we restructure the sequence into different generic sequences based on "meta codons" no longer triplets of 3 nucleotides but values ranging from 17 to 377 nucleotides, then 360 simulations. This method of analysis will then reveal, in most cases, discrete waves or interferences, most often dissonances. However, sometimes there will emerge kinds of resonances where all scales of analysis appear to be in symbiosis.
The discrete interferences fields resulting from the analysis of an entire chromosome are therefore a three- dimensional space:
Dim y (vertical) restructuring in meta codons of lengths 17 to 377 nucleotides Dim x (horizontal) derived mobile1 such that 1/2 1/3 1/4 ... 1 / n
Dim z cumulated populations from the "Master code" operators 14.
The + 1 / -1 derivatives will be of type increase, ie +1 if derivative increasing and will be of type decrease, ie
-1 if derived decreasing.
In this context we will explore these 3d spaces in 2 forms:
-Horizontal, meta codons dimension: curves for a given meta codon dimension, see in the example "resonances" below (see Figure 4).
-Vertically, spectral differentiation: discrete series d2-d1 is +1 if increase and -1 if decrease (see Figure 2 and Figure 3 below).
We represent in top the +1 and in low the -1, (see the examples below).
Example of three-dimensional interference fields (Neanderthal chromosome 1q21.1 DUF1220 rich region6).
Horizontal scan example : 5318499 377103 401874 416639 424005 …/...
(see Figure 4)
Vertical scan example : 1 if d2>d1 and -1 if d2<d1 then : 1 1 -1 1 -1 1 -1 -1 …/...
Figure 2.zoom on vertical scan method from DUF1220 rich region6 Neanderthal 1q21.1.
Figure 3.zoom on vertical scan method revealing main Period = 12 from DUF1220 rich region6 Neanderthal 1q21.1.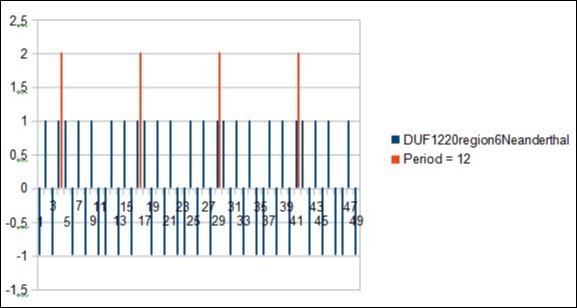
These two independent methods lead in all the cases analyzed to the same period value: here, for example, the main period "horizontal scan" is a resonance of 12bp (Figure 3) and the first 3 "vertical scan" periods are the periods 7 and 12 (Figure 4).
Figure 4.zoom on horizontal scan method revealing Resonances Periods = 5 7 12 from DUF1220 rich region6 Neanderthal 1q21.1.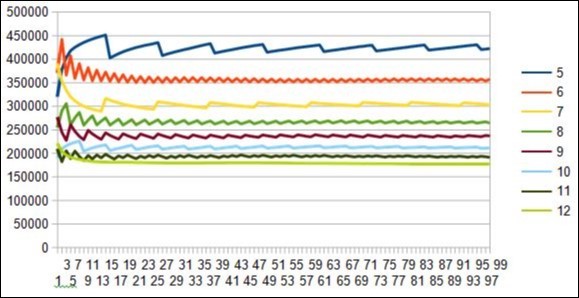
Figure 5.Split of the 5 7 12 19... spectrum in the interference between 2 Fibonacci/Lucas sub-spectrums.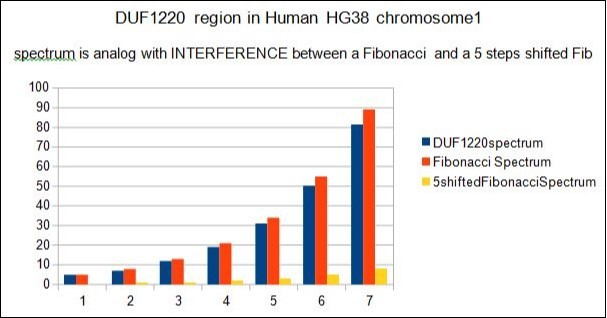
Results
The study of the long region6 of more than 5 million base pairs and containing 218 DUF1220 will reveal the spectrum of the following periods:
First remark: there are various possible interferences between Fibonacci/Lucas sub-spectrums : Main resonances periods:
5 7 12 19 31 50 81 DUF1220 resonances and periods
5 8 13 21 34 55 89 Fibonacci
0 1 1 2 3 5 8 Fibonacci
5 7 12 19 31 50 81 DUF1220 resonances and periods
2 1 3 4 7 11 18 29 47 Lucas
3 6 9 15 24 39 63... = 3 x ( 1 2 3 5 8 13..) = 3 times Fibonacci
5 7 12 19 31 50 81
3 4 7 11 Lucas
4 4 8 12 20 = 4 x (1 1 2 3 5...) = 4 times Fibonacci
5 7 12 19 31 50 81
3 4 7 11 18 Lucas
3 5 8 13 Fibonacci etc...
Second remark : There are main resonances periods like 5 7 12 19 31 50... but also secondary resonances
periods like : 17 (5+12), 24 (5+19), 26 (7+19), 57 (7+50), 69 (50+19), etc...
Main resonances :
5 7 12 19 31 50 81 …/...
Harmonic resonances :
17 24 26 36 38 43 48 55 62 74 91 93 98 …/...
Finally, we could propose the following rule :
The rule is :
"The distance between the waves flow periods sequence from DUF1220 region (5 7 12 19 31 50...) and a
Fibonacci similar sequence (5 8 13 21 34 55...) is ALSO another shifted Fibonacci sequence (0 1 1 2 3 5...) !"
A corollary is:
The waves spectrum associated with DUF1220 region (5 7 12 19... ) is analog with the INTERFERENCE substraction between TWO Fibonacci waves spectrum shifted (then 5 8 13 21 34... and 0 1 1 2 3 5 8 13 ...)
We now study the 16 cases of NBPF genes containing DUF1220 proteins in Homo-Sapiens (HG38), which we will then complete with 3 other representative analogues in Neanderthal.
Recall:
Lucas: 2 1 3 7 11 18 29 47 ...
Fibonacci: 0 1 1 2 3 5 8 13 21 34 …
Finally, we study the 19 cases of DUF1220 regions, including: the six regions from chromosome1 in Homo-Sapiens, then we concentrate on the long region6 of 1q21.1 in Neanderthal, then in the large apes primates, then in Other primates, and finally in other mammals of which the very small number of DUF1220 is known (see Table 1).
Recall :
Lucas 2 1 3 7 11 18 29 47
Fibonacci 0 1 1 2 3 5 8 13 21 34
Here are the details of the periods, resonances and dissonances for the 2 long regions6 of 218 DUF1220 in Sapiens (HG38) and in Neanderthal.
REGION6 sapiens HG38 vs Neanderthal Recall Resonances :
Main resonances : 5 7 12 19 31 50 81 …/...
Harmonic resonances : 17 24 26 36 38 43 48 55 62 69 74 91 93 98 100…/.
In Figure 6, Figure 7, Figure 8, Figure 9, Figure 10, Figure 11 and Figure 12 we illustrate the main resonances and most harmonic resonances. Each of these periods is highlighted by comparing it with the two neighboring numbers, which, in turn, produce dissonances: for example: 18 19 20.
Figure 6.NEANDERTHAL - In this figure we could locate all main periods 7 12 19 31...
Figure 7.SAPIENS HG38 - In this figure we could locate all main periods 7 12 19 31...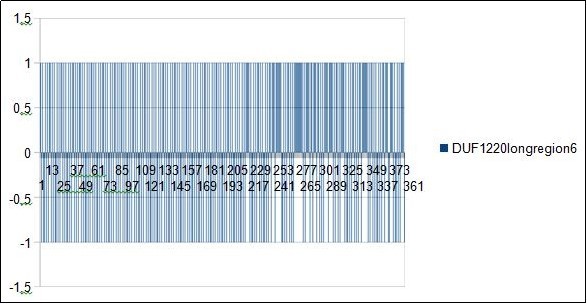
Figure 8.SAPIENS HG38 - In this figure we could locate RESONANCES for all main periods 5 7 12 19 31 50 81
Figure 9.SAPIENS HG38 - In this figure we could locate DISSONANCES shifting by 1 period all main periods 5 7 12 19 31 50 81
Figure 10.NEANDERTHAL - In this figure we could locate RESONANCES for all main periods 5 7 12 19 31 50 81
Figure 11.NEANDERTHAL - In this figure we could locate DISSONANCES shifting by 1 period all main periods 5 7 12 19 31 50 81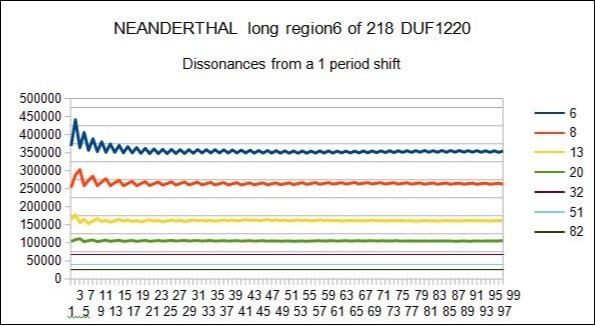
Figure 12.SAPIENS HG38 - This figure show RESONANCES for all harmonic periods 17 24 26 36 38...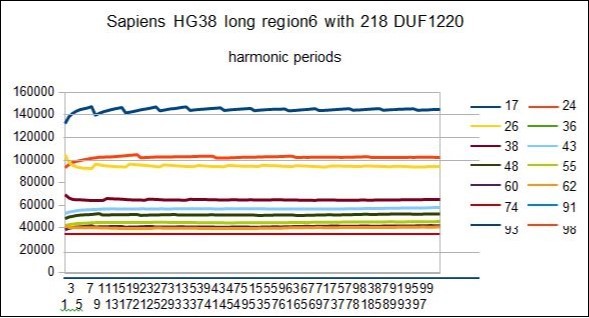
Figure 13.NEANDERTHAL - This figure show RESONANCES for all harmonic periods 17 24 26 36 38...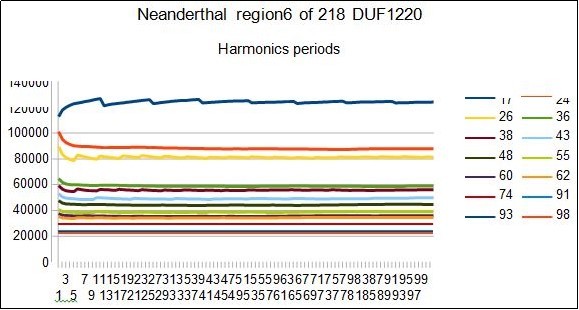
Although Sapiens 17 has a much higher DNA sequencing quality than Neanderthal 16, it should be noted that the results for Neanderthal appear to be more compact than Sapiens. This may be due to the fact that the Neanderthal genomic sample comes from a single (or very small) number of individuals. On the contrary, the genome of Sapiens is a kind of consensus genome coming from a large number of individuals.
Discussion
Illustrated by Figure 14, Figure 15, Figure 16, Figure 17, Figure 18, Figure 19, Figure 20, Figure 21, Figure 22, Figure 23, Figure 24, Figure 25, Figure 26, Figure 27We will discuss more precisely some remarkable points of this digital exploration of the regions DUF1220: Convergence, hierarchization and unification around the numbers of Fibonacci and Lucas of the 4 classes of digital spectra of the regions DUF1220 and the NBPF genes.
The fourfold correlation between different mammalian species, brain evolution, populations of DUF1220 and our numerical classification.
The study of the protein DUF1220.
Dynamic analysis of the long region of 218 DUF1220.
-2- Convergence, hierarchization and unification around the numbers of Fibonacci and Lucas of the 4 classes of digital spectra of DUF1220 regions and NBPF genes.
The study of 19 cases of NBPF genes on the one hand and of 19 regions DUF1220 on the other hand, 38 distinct analyzes appear to us UNIFIED around a single common generic law that can be stated thus:
A / The analysis of each of the regions containing DUF1220 proteins is characterized by a spectrum of numerical periods consisting of a sequence of integers.
B / These numerical spectra are grouped in only 4 characteristic classes.
C / each of these numerical classes can always be written in the form of a simple linear combination between 2 sequences of Fibonacci or Lucas.
D / It emerges a hierarchy allowing to classify these 4 classes among themselves.
Consequently, we can affirm that all the regions containing DUF1220 proteins can be CLASSIFIED, HIERARCHISEE and UNIFIED around a single law linking the numerical spectra of these regions DUF1220 and the sequences of Fibonacci and Lucas.
Let us now go over the rules outlined above.
The table below summarizes the properties of each of these 4 classes.
Let us recall the values of the two suites of Fibonacci and Lucas:
Fibonacci: 0 1 1 2 3 5 8 13 21 34 ...
Lucas: 2 1 3 4 7 11 18 29 47 …
About the HIERARCHY of the 4 classes:
First, we note this remarkable fact: each of the 4 digital spectra always starts with a sequence of 2 consecutive odd numbers (example 5 7).
In the second place, these 4 sequences of odd numbers can be hierarchized with respect to the ratio Phi (where Phi is the number of gold = 1.618033 ...).
5 ÷ 3 = 1.666666667 7 ÷ 5 = 1.4 9 ÷ 7 = 1.285714286 11 ÷ 9 = 1.222222222 13 ÷ 11 = 1.181818182
It will be noted that the best ratio would be the ratio 3/5, where 3 and 5 are numbers of Fibonacci. Although we have not met this ratio among the 38 cases studied, we will specify that the analysis of the large hybrid region fusing the 2 region1 and region2 in 1p has indeed led to a Fibonacci type spectrum, 5 8 13 ... See the figures below.
case 2 :
5 7 12 19 31 50 81 DUF1220 resonances and periods interference between :
2 1 3 4 7 11 18 29 47 Lucas
3 6 9 15 24 39 63... = 3 x ( 1 2 3 5 8 13...) = 3 times Fibonacci
case 3 :
5 7 12 19 31 50 81 DUF1220 resonances and periods interference between :
1 3 4 7 11 Lucas
4 8 12 20 = 2 x 1 1 2 3 5 = 2 times Fibonacci
case 4 :
7 12 19 31 50 81 DUF1220 resonances and periods interference between : 3 4 7 11 18 = Lucas 2 3 5 8 13 = Fibonacci …/.. -2- The fourfold correlation between different mammalian species, brain evolution, populations of DUF1220 and our numerical classification.
Weighting of the 4 classes A B C D: (please see Figure 16 and Figure 17):
The distance of the successive ratios 7/5 9/7 11/9 and 13/11 is calculated to the ideal ratio of 1 / Phi = 0.6180339887. We thus obtain: 0.7142857143 0.7777777778 0.8181818182 0.8461538462
Then:
0.09625172555 0.1597437891 0.2001478295 0.2281198575
Of which we calculate the inverse: 1 / 0.09625172555 0.1597437891 0.2001478295 0.2281198575
= 10.38942413 6.26002429 4.996306992 4.383660462
Which we relativize in order to compare these values to the order of magnitude of the numbers of DUF1220 and the numbers of NBTF.
Finally, by weighting by a coefficient = 10:
10 × 10.38942413 6.26002429 4.996306992 4.383660462 = 103.8942413 62.6002429 49.96306992 43.83660462
Are finally: A: 104 B: 63 C: 50 D: 44
-3- The study of the protein DUF1220.
In Figure 18, Figure 19, Figure 20, Figure 21, and Figure 22, we are interested in the DUF1220 protein itself as well as the gaps separating the 218 successive occurrences in the long region6 of 1q21.1.Here is the sequence of the first of the 118 occurrences DUF1220 of the long region6 of 1q21.1: chr1:144,423,094-144,423,901 808 bp
We analyzed this DNA sequence using the "Master Code of DNA" method described in (15). Here is a summary of the main results:
================== Strand2 =========================================== ANALYSING GLOBAL GENOMICS/PROTEOMICS COUPLING RATIO PERCENTAGE...
UNIVERSALIS GENETIC CODE (Global Unified Genetic Code) GENONOMICS/PROTEOMICS Global Coupling percentage: 90.12953 <===(a)
=== Strand 2 ================================
MAJOR GENOMICS SITES…
15 FIRST + Major sites: 59 60 61 58 62 63 64 51 56 57 65 69 43 55 67<==== (b)
15 LAST - Minorsites: 218 220 243 219 215 222 214 225 221 217 209 201 244 216 208
MAJOR PROTEOMICS SITES...
15 FIRST Major sites: 38 39 40 34 35 36 37 33 44 58 59 41 42 43 32<==== (c)
15 LAST Minor sites: 201 200 199 203 202 240 239 219 218 89 236 235 185 180 87
================= VECTPIM2…
GENOMICS teeth of saw PERCENT CODE...
INCREASE /.\ DECREASE: 0.6230158730.376984127 <==== (d) PROTEOMICS teeth of saw PERCENT CODE...
INCREASE /.\ DECREASE: 0.6111111111 0.3888888889
===================================
PERIODS ANALYSER SUMMARY:
TOPS. PERIODS: 3 7 11 15 19 23 27 31 35 43 47 51 55 59 63 67 71 74 79 83 85 95 …/...<==== €
PERIODS: 4 4 4 4 4 4 4 4 8 4 4 4 4 4 4 4 3 5 4 2 10 5 3 4 4 5 4 3 <==== (f)
AVERAGE PERIODS: 4.285714286
GAUSS PERIODS: VALUE / NB TIMES <==== (g)
Suggested PERIOD: 4
The three figures below and remarks (a) to (d) will enable the reader to grasp the potential of this method whose two strong points are unification between the genomics and proteomics images and On the other hand, a characterization of the fractal texture of the sequence and the demonstration of PERIODS. It is these periods which constitute the elementary brick of our spectral analysis, the object of this article.
Figure 14.example of Fibonacci spectrum related to the mixed region1 and region2 : here 3 5 Resonances.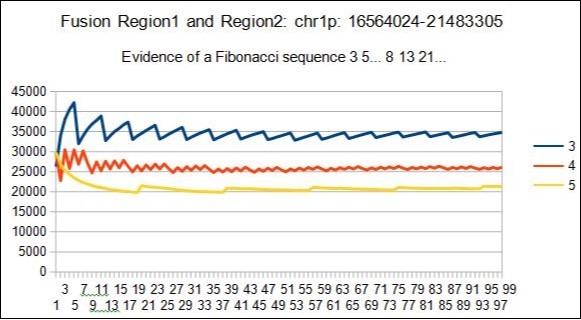
Figure 15.example of Fibonacci spectrum related to the mixed region1 and region2 : here 8 Resonance.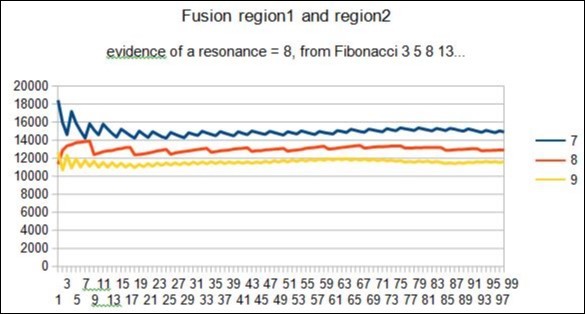
Figure 16.Comparing graphical correlations between DUF1220 numbers and computed normalized spectral classes.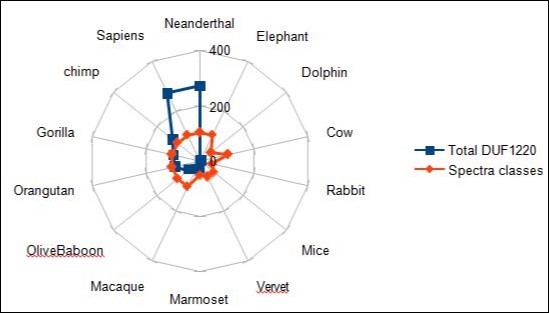
Figure 17.Comparing graphical correlations between NBPF genes numbers and computed normalized spectral classes.
Figure 18.Master Code unifying Genomics and Proteomics patterns for the first DUF1220 protein from the long region6 of 218 DUF1220.
Figure 19.Master Code waves amplitudes for the first DUF1220 protein from the long region6.
Figure 20.Master Code waves fractal textures for the first DUF1220 protein from the long region6.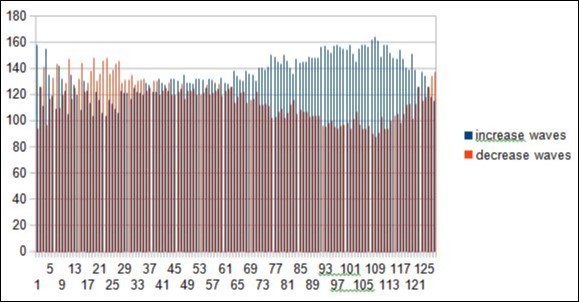
Figure 21.Master Code unifying Genomics and Proteomics patterns for the second GAP saparating DUF1220 proteins from the long region6 of 218 DUF1220.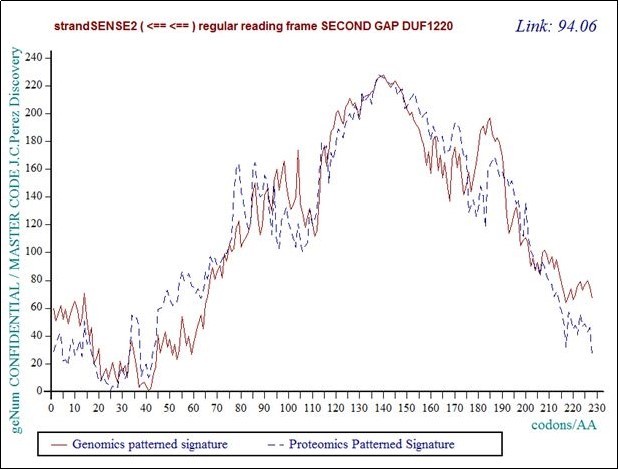
Figure 22.Master Code unifying Genomics and Proteomics patterns for the third GAP separating DUF1220 proteins from the long region6 of 218 DUF1220.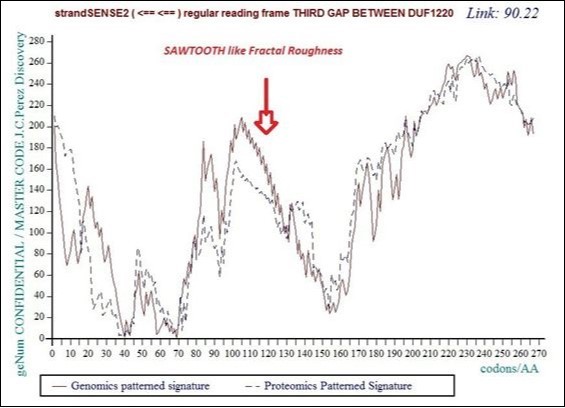
Figure 23.« bar-codes » likes related to the 5 7 12 19 31 spectrum of the 10 first DUF1220 proteins within the long region6 of 218 DUF1220.
Figure 24.Evidence of 5 7 resonances periods in the 5 7 12 19 31 spectrum of the 10 first DUF1220 proteins within the long region6 of 218 DUF1220.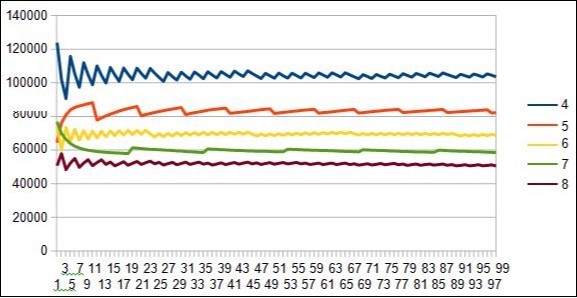
Figure 25.Evidence of 12 resonance period in the 5 7 12 19 31 spectrum of the 10 first DUF1220 proteins within the long region6 of 218 DUF1220.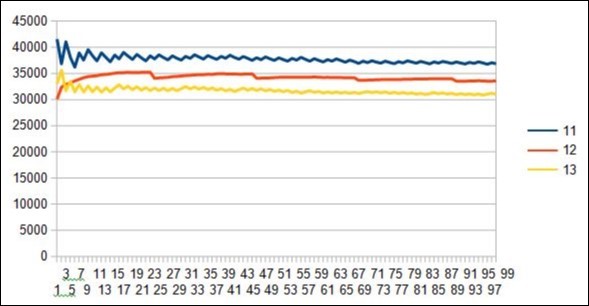
Figure 26.Evidence of 19 resonance period in the 5 7 12 19 31 spectrum of the 10 first DUF1220 proteins within the long region6 of 218 DUF1220.
Figure 27.Evidence of 31 resonance period in the 5 7 12 19 31 spectrum of the 10 first DUF1220 proteins within the long region6 of 218 DUF1220.
Figure 28.Electrical Resonances ( https://en.m.wikipedia.org/wiki/Electrical_resonance ) and Oscilltors like « fork- tuning » vs Nikola Tesla...
Figure 29.the 5 7 12 19 31 spectrum unifying the whole SAPIENS (BUILD34) chromosome10.
Comments:
a) As shown in the figure above, one observes that - for one of the 3 codon reading frames and for one of the reading directions of the sequence - there is a very strong correlation (> 90%) Between the master code image of the double-stranded DNA sequence and the master code of the virtual (or potential) amino acid translation of this same double strand. Here it may seem natural since it is a protein (DUF1220) but the same phenomena are universal throughout the genome, even for non-coding regions! This means that the master code produces a UNIFICATION between the 2 languages of the living being DNA and amino acids. In 15, we show that the third language, the RNA, plays the role of a kind of "neutral element", like the "zero" of mathematics. See the figure opposite (https://www.omicsonline.org/articles-images/2153-0637-5-131- g002.html extracted from Figure 2 in 15.
b) c) It can be seen here that the extreme sites (lowest and highest in the figure) most often have local couplings on the scale of their address of triplets codons. For proteins, we have established that this corresponds to "active sites". For chromosomes, we have established that this corresponds to the points of fragility (breakpoints) of the chromosomes.
d) The "sawtooth" textures (see visual example in Figure 22) are counted here, differentiating the increase and the decreasing slope. We observe that this ratio is very close to 1 / Phi (Phi = 1.618 the golden ratio). As shown in the image on the left, https://www.omicsonline.org/articles-images/2153-0637-5-131-g003.html extracted from Figure 3 in 15 This ratio behaves as an attractor for genomic images of the master code of DNA.
e) f) g) These numerical values illustrate the periodic phenomena of fractal roughness of the textures of the genomics images of the master code. Here we see the sawtooth peaks (e), the corresponding periods (f), and the Gaussian distribution of these periods (g). The probable period is here = 4. Let 4 triplets codons, or ... 3x4 = 12 nucleotides ... This value 12 is precisely the one that we also found in the analysis of the resonancesandperiodsofthisgreatregion6of218DUF1220!
We then wanted to analyze in the same way the "gap" separating 2 consecutive proteins DUF1220.
For example, DUF1220 is the first protein of 218 DUF1220 and INTERDUF1220, the gap separating this first DUF1220 from the second DUF1220:
DUF1220 at chr1:144423094-144423901 start
DUF1220 at chr1:144424700-144425551
INTERDUF1220 in : chr1 : 144423902-144424699
chr1:144,423,094-144,423,901 808 bp.
DUF1220
chr1:144423094-144423901 INTERDUF1220
chr1 : 144423902-144424699
Here their respective compositions in bases T C A G: DUF1220: 179 T, 292 A, 149 T, 188 A.
INTERDUF1220: 179 T, 292 A, 149 T, 188 A.
A detailed analysis will then make us discover this strange fact: the gap separating the first 2 proteins DUF1220 of the sequence region6 of 218 DUF1220 is, itself, a new protein DUF1220! Or, more precisely, the exact sequence capable of coding for a protein DUF1220, because, in fact, just in front of the gene of a protein is most often found a so-called "promoter" sequence ...
The consequence is that a more exhaustive analysis of this region of 218 DUF1220 may expose - perhaps - a number even greater than 218 proteins DUF1220! At the start of the long region6, there are therefore 3 occurrences of the protein DUF1220 attached and consecutive. This long region6 would therefore include not 218, but at least 219 DUF1220 proteins. The entirety of this region must be studied in more detail6. It will also be necessary and above all to carry out this same type of analyses on individual genomes!
Incidentally, the ratios TA and CG at DUF1220 and INTERDUF1220 also show that:
Accumulation DUF1220 = TA = 471
Cumulative DUF1220 = CG = 337
That is, 337 ÷ 471 = 0.7154989384 = 2 - "e".
Where "e" is the Euler constant ( https://en.wikipedia.org/wiki/E_(mathematical_constant ) e = 2.71828.
Indeed, the error is very low: 0.7154989384 ÷ 0.71828 = 0.9961281651
We then carried out the same analyzes on the following 2 gaps. It appears then that they are, on the one hand different from DUF1220 and, on the other hand different from each other.
INTER2DUF1220 second gap
| PERIOD......... | 4 | 19 |
| PERIOD......... | 3 | 3 |
| PERIOD......... | 5 | 3 |
| PERIOD......... | 2 | 1 |
| PERIOD......... | 8 | 1 |
| PERIOD......... | 10 | 1 |
DUF1220 at chr1:144424700-144425551
DUF1220 at chr1:144426288-144427081
144425552-144426287
>hg38_dna range=chr1:144425552-144426287 5'pad=0 3'pad=0 strand=+ repeatMasking=none
INTER3DUF1220
DUF1220 at chr1:144426288-144427081
DUF1220 at chr1:144427831-144428634
144427080-144427830
>hg38_dna range=chr1:144424701-144425552 5'pad=0 3'pad=0 strand=+ repeatMasking=none
In this figure the reader observes the "saw teeth" typical of the phenomenon of fractal roughness, at the origin of the periods and resonances.
-4- Dynamic analysis of the long region of 218 DUF1220
In Figure 23, Figure 24, Figure 25, Figure 26, Figure 27, we will now consider a possible dynamic, perhaps "threshold effects" that could result from the number of successive DUF1220. For this, in the long region of 218 DUF1220, we have successively analyzed the periodic spectra:
Of the first 5 DUF1220,
Of the first 10 DUF1220,
Of the first 20 DUF1220,
-of the first 50 DUF1220,
Here is an overview of these results, the reader will find the complete study in complementary materials (Part IV). Reminder of the sequence of 218 DUF1220:
Chr1q
Case2 : Analyse des 10 premiers DUF1220 :
DUF1220 at chr1:144423094-144423901 start
DUF1220 at chr1:144424700-144425551 DUF1220 at chr1:144426288-144427081 DUF1220 at chr1:144427831-144428634
DUF1220 at chr1:144429713-144433807
DUF1220 at chr1:144435163-144435809
DUF1220 at chr1:145291607-145336848
DUF1220 at chr1:145293214-145294055
DUF1220 at chr1:145294792-145295601
DUF1220 at chr1:145296351-145297173
1: 144423094-145297173
DUF1220by10
Chromosome 1: 144,423,094-145,297,173
Conclusion:
As illustrated by Figure 28, in this long “contig” (DNA region) of more than 5 million bases of the region 1q21.1, comprising 218 DUF1220, What does this potential of main resonances mean?
Main resonances periods:
5 7 12 19 31 50 81
DUF1220 resonances and periods
+ 0 1 1 2 3 5 8 Fibonacci
= 5 8 13 21 34 55 89 Fibonacci
5 7 12 19 31 50 81
DUF1220 resonances and periods
- 2 1 3 4 7 11 18 29 47 Lucas
= 3 6 9 15 24 39 63... = 3 x ( 1 2 3 5 8 13..) = 3 times Fibonacci
This means, according to our intuition, that this region of spectrum (5 7 12 19 31 50 81) represents the first part of a kind of potential "resonator".
Then, "interference" with an external spectrum of Fibonacci in addition (+ 0 1 1 2 3 5 8) or with a Lucas spectrum in subtraction (-2 1 3 4 7 11 18 29 47) will produce a "resonance "Of type Fibonacci (= 5 8 13 21 34 55 89) or else (3 x 1 2 3 5 8 13 ..) ...
Certainly, this second part of the resonator coming from an external source would remain to be determined ... Thus, in a kind of "key-lock" system, this 1q21.1 region would then act as a kind of "receiver", the "lock"!
But a receiver of what type "transmitter", the "key"?
The profoundly bipolar character of this phenomenon remains indivisible.
Can one imagine how the perception of the image of a nautilus or the musical resonance of Bach or the subtle aesthetic of a mathematical equation could constitute the beginning of the process constituting this "key"?
Certainly the genome when studied as a mathematical object with the tools of mathematical topology can offer a medium of resonance via the subtle topologies of the ribbon of Moebius or the bottle of Klein as demonstrated by the studies of Diego Lucio Rapoport 18.
Indeed, as demonstrated by Volkmar Weiss 19, the proportions of Phi and the number of gold in the electro-encephalogram (EEG) are discovered on the scale of experimental measurement.
During our Artificial Intelligence research at IBM, based on our model "Fractal Chaos" 20, 21, 22, we had demonstrated the numerical hyper-sensitivity of an artificial neural network positioned in the vicinity of Phi and Subjected to perturbations of the Fibonacci number type 23, 24.
The fractal architecture 25, 26, the mathematics 27 and Phi the golden number are omnipresent in Nature, in humans, in DNA, and even on the atomic scale 28 ... It is a serious mistake to fall into these simplistic theses of an omnipresence of the golden number in the DNA as well as an omnipresence of waves in the DNA ...
The reality that we have discovered for more than 25 years of biomathematics exploration of DNA is much more subtle:
Phi the Golden ratio 29, 30, 31, 32, 33 and the numbers of Fibonacci never appear in the same form according to the scale and level of genetic information studied:
In 1991 11 we discover proportions of Fibonacci structuring the proportions of bases TCAG inside the sequences of the genes ... but not in the junk DNA non coding.
In 1997 14, 15 we discovered that a numerical law of projection of the atomic masses of the base and amine acids, a law based on pi and phi, unifies the 3 languages of the genetics that are DNA, RNA, amino acids. In 2009 34, 35 we demonstrate that the fractal texture of the above unified genomic and proteomic images is manifested in the form of periodic waves.
Finally, from 2010 36, 37 we publish various articles demonstrating that the golden ratio controls and finely adjusts the proportions of triplets codons to the scale of the entire human genome.
This multi-level diversity of Phi is as astonishing as it is radiant ...
For example, in front of the cluster formed by the four discoveries presented in this article on the one hand, in 38 on the other hand, and finally in 39 we can only remain remarkable:
In this paper, we show that there is a sort of HGO (human genome optimum) unifying the entire human genome. This HGO will then allow the discovery of a universal law guiding all the LOH deletions implied in the cancers.
In "Humans and Primates Chromosomes4 Fractal CODES: periodic stationary waveforms characterizing and differentiating Neanderthal and Sapiens whole chromosomes DNA sequences" 39 we show a sort of hierarchy classifying the 24 human chromosomes. One of them, the chromosome4 would be a kind of leader, "lighthouse" ... and this lighthouse vibrates to the proportions of Phi, the numbers of Lucas and Fibonacci.
Finally, in this article we discover more strange still: Phi and the numbers of Fibonacci and Lucas no longer appear explicitly. They hide behind a subtle interference, hiding behind a spectrum of numerical periods in the appearance of any numbers.
Our intuition is that there exists a link, "dialogues" between these three heterogeneous but subtle mechanisms ...
We suggest in this scenario Key lock or transmitter receiver, a probable link between the chromosome4 of which 39 will show that it behaves like a kind of resonance emitter of Fibonacci and Lucas ...
And this famous region 1p21.1 with more than 200 DUF1220 would then behave like a kind of "resonance box", the resonator.
Finally, in "Global and long range fractal differences between sapiens and Neanderthal genomes" 40, we will demonstrate how the law discovered here seems to apply sometimes even to the scale of an entire chromosome such as chromosomes 10 and 11, The image of chromosome 10 of Sapiens BUILD34 below, controlled by a spectrum 12 19 31 ...
Multidisciplinary considerations
If this article had been an article of “pure mathematics” we could have discussed the convergence properties of the respective sequences discovered here:
1 3 4 ... (Lucas)
3 5 8 ... (Fibonacci)
5 7 12 ...
7 9 16 ...
9 11 20 ...
11 13 24 ...
... / ...
But this article concerns genetics and human genomics ... The genetics of the densest protein region within the whole human genome: DUF1220 protein region.
The genomics of the longest of our chromosomes, the chromosome 1.
How can we explain that these sequences of DUF1220 are organized in periodic structures obeying these numerical sequences ALL constructed from 2 consecutive odd numbers?
How to explain this total correlation between the levels of evolution of the 4 classes of sequences observed (Table 4, Table 5 and Table 6) on the one hand and the levels of evolution of different mammals on the other?
Table 4. The spectrums related to 19 cases of DUF1220 regions from Sapiens, Neanderthal, primates and other mammals reveals 4 classes of spectrums, all could be split in Fibonacci/Lucas sub-spectrums.| DUF1220 regions | Number of DUF1220 | Resonance Periods | Fibonacci/Lucas Periods | Shifted sequences | Hyper - resonance class | |
| Anthropoids Primates | ||||||
| Region11:16564024 16587759 | 7 | 9 11 20 31 51 | 8 8 16 24 40 = 8x(1 1 2 3 5) | 1 3 47 11 | C | |
| Region21:21472881 21483305 | 5 | 7 9 16 25 41 | 5 8 13 21 34 | 2 1 3 4 7 | B | |
| Region31:108232266 108239862 | 4 | 7 9 16 25 41 | 5 8 13 21 34 | 2 1 3 4 7 | B | |
| Region4108454483 108462078 | 4 | 7 9 16 25 41 | 5 8 13 21 34 | 2 1 3 4 7 | B | |
| Region51:120809849 120840512 | 7 | 7 9 16 25 41 | 5 8 13 21 34 | 2 1 3 4 7 | B | |
| Region61:144423094- 149554657 | 218 | 5 7 12 19 31 50 81 | 5 8 13 21 34 55 89 | 0 1 1 2 3 5 8 | A | |
| Region6 | 5 7 12 19 31 50 81 | 5 8 13 21 34 55 89 | 0 1 1 2 3 5 8 | A | ||
| NEANDERTHAL1:141527445-146659008 | ||||||
| Chimp region61:111091664-116223227 | 5 7 12 19 31 50 81 | 5 8 13 21 34 55 89 | 0 1 1 2 3 5 8 | A | ||
| Gorilla region61:124329600-129461163 | 5 7 12 19 31 50 81 | 5 8 13 21 34 55 89 | 0 1 1 2 3 5 8 | A | ||
| Orangutan region61:102601348-107732911 | 5 7 12 19 31 50 81 | 5 8 13 21 34 55 89 | 0 1 1 2 3 5 8 | A | ||
| Olive Baboon Region61:119609055-124740618 | 5 7 12 19 31 | 8 13 21 34 | 1 1 2 3 | A | ||
| Macaque region61:120005413-125136976 | 5 7 12 19 31 50 81 | 5 8 13 21 34 55 89 | 0 1 1 2 3 5 8 | A | ||
| Marmoset region618:1-5131564 | 9 11 20 31 51 | 8 8 16 24 40 = 8x(1 1 2 3 5) | 1 3 47 11 | C | ||
| Vervet Region61:1-1280666 | 7 9 16 25 41 | 5 8 13 21 34 | 2 1 3 4 7 | B | ||
| Prosimians | ||||||
| Mice Region63:94635287-99766850 | 7 9 16 25 41 | 5 8 13 21 34 | 2 1 3 4 7 | B | ||
| Rabbit Region613:40672536-45804099 | 11 13 24 37 | 2 1 3 4 | 9 12 21 333 x (3 4 711) | D | ||
| Laurasiatheria | ||||||
| Cow Region63:19523691-24655254 | 5 7 12 19 31 50 81 | 5 8 13 21 34 55 89 | 0 1 1 2 3 5 8 | A | ||
| Dolphin region6GeneScaffold_2231:20869 5:327249 | 9 11 20 31 51 | 8 8 16 24 40 = 8x(1 1 2 3 5) | 1 3 47 11 | C | ||
| Afrotheria | ||||||
| Elephant region6scaffold_127:1-2319825 | 5 7 12 19 31 50 81 | 5 8 13 21 34 55 89 | 0 1 1 2 3 5 8 | A | ||
| Class reference | Hierarchy | Basic initial fractal periods spectrum... ==> | ==> Interfering with Fibonacci or Lucas pattern ... ==> | ==> Result of the numerical interference... reveals another Fibonacci or Lucas pattern | Number of DUF1220cases |
| A | High = 1 | 5 7 12 19 31 50 81 | + 0 1 1 2 3 5 8 | = 5 8 13 21 34 55 89 | 12 |
| B | Medium = 2 | 7 9 16 25 41 | - 2 1 3 4 7 | = 5 8 13 21 34 | 16 |
| C | Medium = 3 | 9 11 20 31 51 | - 1 3 47 11 | = 8 8 16 24 40 = 8x(1 1 2 3 5) | 9 |
| D | Low = 4 | 11 13 24 37 | - 2 1 3 4 | = 9 12 21 33 = 3x(3 4 7 11) | 1 |
| Species | Total DUF1220 | Spectra class | NBPF genes |
| Neanderthal | 272 | 104 | 23 |
| Sapiens | 272 | 104 | 23 |
| chimp | 125 | 104 | 15 |
| Gorilla | 99 | 104 | 11 |
| Orangutan | 92 | 104 | 10 |
| OliveBaboon | 53 | 104 | 10 |
| Macaque | 35 | 104 | 10 |
| Marmoset | 31 | 50 | 11 |
| Vervet | 3 | 63 | 2 |
| Mice | 1 | 63 | 0 |
| Rabbit | 8 | 44 | 3 |
| Cow | 7 | 104 | 3 |
| Dolphin | 4 | 50 | 1 |
| Elephant | 1 | 104 | 2 |
Acknowledgements
We especially thank Dr. Volkmar Weiss who suggested we look for possible hidden codes in these long 1q21.1 DNA sequences containing large DUF1220 proteins population. We also thank the mathematician Diego Lucio Rapoport (Buenos aires), the biologist François Gros (Pasteur Institut,co-discoverer of RNA messenger with james Watson and walter Gilbert ) and Professor Luc Montagnier, medicine Nobel prizewinner for their interest in my research of biomathematics laws of genomes.
References
- 1.J M Sikela. (2006) The jewels of our genome: the search for the genomic changes underlying the evolutionarily unique capacities of the human brain. PLoS Genetics.
- 2.J M Davis, Quick Searles, B V, J M Sikela. (2015) Replicated linear association between DUF1220 copy number and severity of social impairment in autism. , Human Genetics 134(2015), 569-575.
- 3.Weiss V. (1992) Major genes of general intelligence. in Personality and Individual Differences, ELSEVIER, Volume 13, Issue 10 1115-1134.
- 4.J M Davis, V B Searles, Anderson N, Keeney J, Raznahan A et al. (2015) DUF1220 is linearly associated with increased cognitive function as measured by total IQ and mathematical aptitude scores. , Human Genetics 134(2015), 67-75.
- 5.Dumas L, J M Sikela. (2009) DUF1220 domains, cognitive disease, and human brain evolution. , Symposia on Quantitative Biology 74, 375-382.
- 6.J G Keeney, Dumas L, Sikele J M Sikela. (2014) The case for DUF1220 domain dosage as a primary contributor to anthropoid brain expansion. Frontiers in Human Neurosciencedoi.org/10.3389/fnhum.2014.00427.
- 7.O’Bleness M, V B Searles, C M Dickens, Astling D, Albracht D et al. (2014) Finished sequence and assembly of the DUF1220-rich 1q21 region using a haploid genome. , BMC Genomics 15, 2164-15.
- 8.Zimmer F, S H Montgomery. (2015) Phylogenetic analysis support a link between DUF1220 domain number and primate brain expansion. , Genome Biology and Evolution 7, 2083-2088.
- 9.Wu Jun. (2016) (2016)In vivogenome editing via CRISPR/Cas9 mediated homology-independent targeted integration,Nature(16. doi: 10.1038/nature20565
- 10.Lieberman-Aiden. (2009) Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science. , DOI:
- 11.J C Perez. (1991) Chaos DNA and neuro-computers: a golden link. Speculations in Science and Technology 14, 336-346.
- 12.P J Marcer. (1992) Order and chaos in DNA – the Denis Guichard Prizewinner: Jean-Claude Perez. , Kybernetes 21, 60-61.
- 15.Perez J. (2015) Deciphering Hidden DNA Meta-Codes -The Great Unification &. , Master Code of Biology. J Glycomics Lipidomics 5, 131-10.
- 18.Diego L Rapoport. (2016) Klein Bottle Logophysics, Self Reference, Heterarchies, Genomic Topologies, Harmonics and Evolution. Part III. The Klein Bottle Logic of Genomics and its Dynamics, Quantum Information, Complexity and Palindromic Repeats in Evolution. Quantum Biosystems. 7(1), 107-174.
- 19.Weiss H, Weiss V. (2003) The golden mean as clock cycle of brain waves, Chaos, Solitons & Fractals.
- 20. (1988) De nouvelles voies vers l'intelligence artificielle: pluri-disciplinarite: auto-organisation: réseaux neuronaux. , JC Perez, Masson Paris
- 22.CHAOS” “FRACTAL.a new neural network holographic model JC Perez. , JM Bertille, Neural Networks 1, 121.
- 23.Perez J C. (1990) Digital holograms computers, concepts and applications in Neural Networks / Les Reseaux de Neurones: Biological Computers or Electronic Brains / Ordinateurs Biologiques Ou Cerveaux Eletroniques. Les Entretiens de Lyon. Proceedings of the International Conference "Les Entretiens de Lyon" on Neural Networks. Ec... (Paperback), Published , ISBN 2-287.
- 24. (1990) Integers neural network systems (INNS) using resonance properties of a Fibonacci's chaoticgolden neuron'. IJCNN International Joint Conference on , JC Perez, Neural Networks 859-865.
- 25.Pellionisz A J, Graham R, Pellionisz P A. (2012) Recursive Genome Function of the Cerebellum: Geometric Unification of Neuroscience and Genomics. In:. Manto M, DL, et al. (Eds) Handbook of the Cerebellum and Cerebellar Disorders. 1st (Edn) .
- 26.Dokukin M E, Guz N V, Woodworth C D, Sokolov I. (2015) Emergence of fractal geometry on the surface of human cervical epithelial cells during progression towards cancer. , New J Phys 17.
- 28.Coldea R, D A Tennant, E M Wheeler, Wawrzynska E, Prabhakaran D et al. (2010) Quantum Criticality in an Ising Chain: Experimental Evidence for Emergent. E8 Symmetry. Science .
- 29.Friedman R, Cross M. (2013) The Golden Ratio & Fibonacci Sequence: Golden Keys to Your Genius, Health. Wealth & Excellence. 1st (Edn) , Hoshin Media, USA .
- 30.Persaud Dharam, James P O'Leary. (2015) Fibonacci Series, Golden Proportions, and the Human Biology. , HWCOM Faculty Publications. Paper 27, 27.
- 33.Vaezi A, Barkeshli M. (2014) Fibonacci Anyons From Abelian Bilayer Quantum Hall States." Physical Review Letters 113.
- 34.S A Olsen. (2017) Riding the Numbers: Ratios & Resonant States of Consciousness. http://www.hrpub.org DOI: 10.13189/sa.2017.050108 , http://www.hrpub.org/download/20161230/SA8-19608211.pdf , Sociology and Anthropology 5(1), 72-75.
- 35.Perez J C. (2011) Caminos Interdisciplinaios. Seminario CLAVE_INTER, Espacio Interdisciplinario, Universidad de la Republica. , Montevideo, Uruguay
- 36.Perez J C. (2011) Decoding Non-Coding DNA Codes: Human Genome Meta-Chromosomes Architecture. BIT Life Sciences’ 3rd Annual World Vaccine Congress , Beijing, China .
- 37.Perez J C. (2010) Codon Populations in Single-Stranded Whole Human Genome. DNA Are Fractal and Fine-Tuned by the Golden Ratio 1.618. Interdisciplinary Sciences: Computational Life Sciences 2, 1-13.
- 38.Perez J C. (2013) The “3 Genomic Numbers” Discovery: How Our Genome Single-Stranded DNA Sequence Is “Self-Designed” as a Numerical Whole. , Applied Mathematics 4, 37-53.
- 39.Jean Claude Perez.The Human Genome Optimum (HGO): Towards a Universal Law Controlling All Human Cancer Chromosome LOH Deletions.
- 40.CP Cancer Sci. (2017) 1[1]: 007Perez J.C., "Humans and Primates Chromosomes4 Fractal CODES: periodic stationnary waveforms charaterizing and differenciating Neanderthal and Sapiens whole chromosomes DNA sequences", to be published.
- 41.J C Perez. (2017) Global and long range fractal differences between sapiens and neanderthal genomes", to be published.
- 42.Perez J C. (2017) Sapiens Mitochondrial DNA Genome Circular Long Range Numerical Meta Structures are Highly Correlated with Cancers and Genetic Diseases mtDNA Mutations. , J Cancer Sci Ther 9, 512-527.
- 43.Roman V Yampolskiy. (2017) On the origin of synthetic life: attribution of output to a particular algorithm. , The Royal Swedish Academy of Sciences , Phys. Scr 92, 10-1088.
- 44.Montagnier L, Aïssa J, Ferris S, Montagnier J L, Lavallée C. (2009) . Electromagnetic Signals Are Produced by Aqueous Nanostructures Derived from Bacterial DNA Sequences. Interdiscip Sci Comput Life Sci 1, 81-90.
Cited by (2)
This article has been cited by 2 scholarly works according to:
Citing Articles:
Jean-claude Perez - (2018) Semantic Scholar
