Molecular and Metabolic Pathogenesis of Familial Combined Hyperlipidemia and Association with Metabolic Syndrome
Abstract
Background
The objective of this review is to unify the various genetic defects along with elaborating metabolic pathways in Familial Combined Hyperlipidemia(FCHL) and also to differentiate the phenotype of FCHL from metabolic syndrome.
Methods
PubMed and Cochrane’s library was searched for keyword “Familial combined hyperlipidemia” and latter with “Familial combined hyperlipidemia genes” to finally shortlist 23 articles. Further search with key words “molecular pathogenesis of familial combined hyperlipidemia” and “metabolic syndrome and familial combined hyperlipidemia” was carried out for finding molecular defects in FCHL, non-molecular findings distinguishing FCHL from metabolic syndrome and overlapping features between FCHL and metabolic syndrome.
Results
Major culprit regions identified included Chromosome-1q21-q24(USF1 and FOXA2) , Ch-11q (APOA5), Ch-16q24, Ch-20q12-q13.1, Ch.4q32.3 (rs6829588), and Ch-19q13.32 containing PVRL-2 gene (Also known as Nectin-2). The genetic and metabolic pathways linked to FCHL may involve: 1-Defective clearance of Apo-B containing lipoproteins, 2-Overproduction of Apo-B containing lipoprotein i.e., VLDL and 3-Adipose tissue dysfunction. FCHL phenotype showed close resemblance with metabolic syndrome clinical and biochemical features with slight differences.
Conclusion
The reviewed data suggested that FCHL phenotype is the resultant end outcome from multiple molecular defects and thus underlying genetic defect identification in the index case is important for personalized medicine and incoming gene therapy. Further research is warranted to explore specific genetic defects.
Author Contributions
Academic Editor: Kentaro Kato, Department of Eco-epidemiology, Institute of Tropical Medicine, Nagasaki University
Checked for plagiarism: Yes
Review by: Single-blind
Copyright © 2019 Sikandar Hayat Khan
 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Competing interests
The authors have declared that no competing interests exist.
Citation:
Introduction
Familial Combined Hyperlipidemia (FCHL) supersedes most other genetic lipid and non-lipid disorders in most of the shared scientific evidence with disease prevalence ranging from 1% to 2.5% in different parts of the globe. 1 Apparently the available data on the subjects suggests the significantly heterogeneous FCHL genetics, extremely high risk of premature coronary artery disease (CAD), hypertriglyceridemia related disorders, and relationship with other metabolic disorders like insulin resistance. 2, 3 Unmet need must also be acknowledged on its impact on health economy in terms of this being the most frequent lipid disorder, loss of productive technical man hours, family crises emerging from premature morbidity and mortality and linkage or lead to newer metabolic disorders. 4
FCHL is not a singular entity in pathology or a monogenic genetic disorder. Molecular and genetic studies and phenotypic disease presentations have identified it as heterogeneous disease which not only vary in terms of temporo-spatial presentation clinical traits, being very pleiotropic due to probable multiple molecular triggers and the clinical appreciation variability resulting from the subtype of cholesterol surge or decline and very high prevalence in myocardial survivors. 5, 3, 6 All these differential presentation surrounding the singular label “FCHL” and inputs from small scale studies focusing upon few aspects of this broad-spectrum pathology makes it difficult to define the disease. 7 Nonetheless literature search converges to a few common traits which may help elucidate a criteria for nominating a person with FCHL. These criteria includes: 1- Strong family history (at least 2 members from one family), 2- Variable lipid abnormalities including combined hyperlipidemia or hypercholesterolemia or hypertriglyceridemia all shaping in different dimension over a person’s life, 3- Autosomal dominant pattern of inheritance with high penetrance, 4- Dermatological manifestations like xanthomas are rare unlike familial hypercholesterolemia and 5- Strong predisposition to early onset coronary artery disease (CAD). 8
Most primary physicians lack the knowledge and details about the gravity underlying the interpretation of a lipid profile along with being oblivious of underlying lipid pathology, so patients are dealt as per conventional wisdom with disregard to any attempt to provide the patient with complete diagnosis along with advice on family screening. Furthermore, the disease is in definitive demand of a biochemical or molecular biomarker to allow predictive screening, identifying the specific genetic defect in lipid physiology, genetic risk scoring, cascade screening of family members, preventive guidance to carriers and patients and most importantly to provide a way forward to curative medicine by gene therapy like CRISPR-Cas9 techniques. 9
This mini review attempts to segregate the molecular subtypes of FCHL, brief details of molecular pathogenesis, overlapping and contrasting evidence linking FCHL and metabolic syndrome in order to bridge the gap with some recent data and not so recent concepts about FCHL.
Review Methodology
The key words “Familial combined hyperlipidemia” was searched on PubMed and Cochrane’s library. Search with initial keyword generated 1058 articles on PubMed which was narrowed down by using specific key words as “Familial combined hyperlipidemia genes” yielded 37 articles. Articles without complete text were excluded, which further reduced the list to 23. Later search with key words “molecular pathogenesis of familial combined hyperlipidemia” and “metabolic syndrome and familial combined hyperlipidemia” to yield 113 and 93 articles which were shortlisted by applying filters initially and reviewed for finding molecular defects in FCHL, non-molecular findings distinguishing FCHL from metabolic syndrome and overlapping features between FCHL and metabolic syndrome.
Molecular and Metabolic Mechanisms Behind FCHL
Though the aforementioned criteria provide a dimension to diagnosis with some degree of sensitivity but in clinical practice it is far from being specific. The problem part starts in clinics where the defining criteria are not equated with clinical presentation of the patient due to variable phenotypic presentations. Attributes to molecular pathogenesis of “FCHL” have been varying across the literature. Most evidence related to date focus on USF1 transcription factor located on Chromosome-1q21-q24. 10, 13 Pajukanta et al pioneered the genetic work by elucidating the region Ch.1q21-24 nearby Apolipoprotein-A2 region through linkage analysis to provide a LOD of 5.93 thus indicating strong phenotype-genotype linkage along with a possibility dominant mode of inheritance for FCHL. 10 These studies in fact led to the initial molecular work shortlisting Ch.1q23.3 as culprit region for FCHL and provided us some molecular pathological details about USF1 gene to combined hyperlipidemia. (Figure 1) However, later studies identified other genetic loci and biochemical evidence to suggest other molecular pathological defects leading to a similar phenotype. de Graaf et al have suggested that the presence of obesity and insulin resistance accompany subjects diagnose to have FCHL with probable defective adipose tissue metabolism leading to increase in free fatty acids could be the main culprit implying a defect in adipose tissue metabolism. 14 Wijsman et al have identified that Apo-B levels and size are both major determinants associated with FCHL with SNP genotyping identifying an important candidate region on Ch.4q32.3. 15 Further studies in the past have identified multiple other loci which could lead to a FCHL phenotype apart from Chromosome: 1q21 (USF1) gene, including candidate genes like Lipoprotein lipase gene and Apo lipoproteins A1/c3/a4/a5 genes. 16 Another study by Cruz-Bautista et al have shown that atherogenic particles like Apo-B containing VLDL, along with Apo-CIII and AII along with adiponectin hormone appear to be the major factors involved in defining FCHL phenotype. 17 Other related evidence is presented in Table 1 and Table 2. Converging these pieces of evidences together, it seems that multiple factors and mutations/polymorphisms at different genetic loci are involved in shaping up the clinical phenotype of “FCHL”.
Figure 1.Genetic loci for USF1 gene, its association and relation with FOXA2 and hepatic triglyceride secretion along with role of SNP rs2073658 in USF1 gene in altering lipid and other metabolic effects.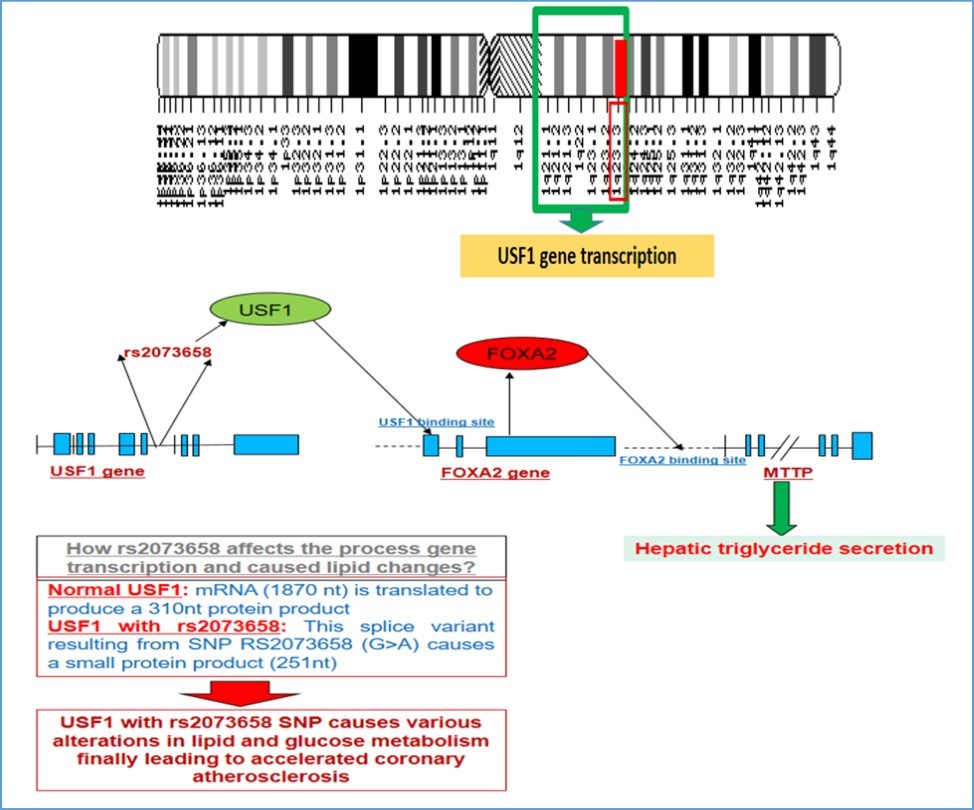
| Ser | Genetic defect | Data summary | Metabolic syndrome association | References |
|---|---|---|---|---|
| 1. | Chromosome-1q21-q24SLC25A40LASS4 | WES and Linkage studies identified role of PCSK9 and phospholipid transfer protein to be involved along with previously known defects in FCHL | Yes | van Greevenbroek et al. (2014) 12 |
| 2 | Chromosome-1q21-q24(USF1 transcription factor )APOC3Ch-11q:APOA5(Linked with hypertriglyceridemia) Ch-16q24 & Ch-20q12-q13.1 | The author contributed upstream factor 1 on C-1 as a major factor linked with FCHL. Apart from involvement of ch-1 multiple other loci with small effects were found which probably cause the phenotypic variability | Yes | Naukkarinen et al. (2006) 14 |
| 3. | LDL-receptor mutationsLipoprotein Lipase mutations | The evaluated clinical phenotypes of FCHL with main focus on LDL-Receptor and Lipoprotein lipase mutations to conclude that some categories can be variants of familial hypercholesterolemia, while the lipoprotein lipase gain of function and Loss of function mutation did not vary significantly between control subjects and cases of FCHL. | Yes | Minicocci et al (2015) 18 |
| 4. | Multiple small impact genetic changes | DNA sequencing study identified multiple small lipid genetic effect changes rather than major impact causing mutation. Resulting rise of either singular lipid type or combined lipid rise. | NA | Brahm et al. (2016) 19 |
| 5. | Lp(a) major contributor towards IHD in mutation negative FCHL | *>50% of phenotypic FCHL cases had no mutation (n=206)*Lp(a) were positive in up to 45% of such cases*CAD association more probable with Lp(a), not FCHL | Yes | Ellis et al. (2016) 20 |
| 6. | Elevated Apo-BFeatures of metabolic syndrome | Acknowledged role of environment along with multiplicity of genetic alterations | Yes | Bello-Chavolla et al (2018) 6 |
| 7. | Ch.4q32.3 (rs6829588) | Linkage analysis by SNP genotyping on Ch.4q.32.3 idefied a strong candidate region (rs6829588) for being associated with 39% of Apo-B variance in FCHL subjects | NA | Wijsman et al(2010) 15 |
| 8. | Up to 35 genes | Overlapping genes with affecting adipose function of adipose tissue, hepatocyte fat deposition, clearance/disposal of Apolipoprotein B lipoproteins were analyzed using candidate genes and genetic association in subjects with FCHL leading to multiple phenotypes | Yes | Brouwers et al. (2012) 2 |
| 9. | Ch-19q13.32 containing PVRL-2 gene (Also known as Nectin-2) | This GWAS has highlighted polymorphism at PVRL-2 region is associated with FCHL like phenotype with components of metabolic syndrome | Yes | Mo et al.(2018) 21 |
| 10 | Ch.1q21-23Ch.11pCh.16q22-24.1 | This linkage-association study identified following candidate genes in FCHL:Lipoprotein lipase (LPL) geneApo-A1/C3/A4/A5 genes Ch.1q21(USF1) gene | Yes | Suviolahtiet al (2006) 16 |
| 11. | Ch.1 (USF1) gene | rs2073658 located in FOXA2 modulates the triglyceride transfer in microsomes by feed forward loop involving USF1 gene activation | Yes | Auer et al (2012) 22 |
| 12 | Ch. 20q12-q13.1 | Hepatic Nuclear Factor-4 alpha (HNF4A) leads to alteration in lipid profile, which overlap between metabolic syndrome and FCHL. | Yes | Weissglas-Volkov et al 23 |
| Ser | Identified metabolic defects | Overlap with metabolic syndrome | References |
| 1 | Identified defects include: Adipose tissue turnover defect, late plasma clearance VLDL & chylomicron, increased VLDL production, , LDL disposal is delayed | Yes | Taghizadeh et al (2019) 24 |
| 2. | FCHL has been associated with components of metabolic syndrome | Yes | Díaz-Ruiz et al. (2019) 25 |
| 3. | PCSK9 levels are associated with FCHL and components of metabolic syndrome | Yes | Vlachopoulos et al. (2018) 26 |
| 4. | FCHL is overlapping with multiple metabolic features and probably have polygenic inheritance | Yes | Vaverková et al. (2018) 27 |
| 5 | Plasmingen Activator Inhibitor-1 and myelperoxides are associated with insulin resistance in subjects with FCHL | Yes | Carratala et al. (2018) 28 |
| 6 | Dyslipidemia type IIb are shared by type-2 diabetes mellitus, metabolic syndrome and FCHL | Yes | Arai et al (2012) 29 |
| 7. | The prevalence of FCH in subjects diagnosed with Metabolic Syndrome is 36.7% | Yes | Pie et al (2007) 30 |
| 8 | Prolong post-prandial hyperlipidemia may be associated with both metabolic syndrome and FCHL | YES | Alipour et al (2007) 31 |
| 9. | Apo-B was found to be prevalent in subject with FCHL | YES | Pei et al (2005) 32 |
The aforementioned referenced discussion over last 3 decades thus concluded “phenotype FCHL” to be emerging from various genetic alterations across various chromosomal regions with slightly different clinical presentations. The genetic and metabolic pathways linked to FCHL may involve: 1-Defective clearance of Apo-B containing lipoproteins, 2-Overproduction of Apo-B containing lipoprotein i.e., VLDL and 3-Adipose tissue dysfunction. A detailed account of various genes affected in the above pathway has earlier been given by Brouwers et al. 2, Figure 2 explains an overview of pathways and key areas proposed to be affected in different organs identifying the multifaceted dimensions to FCHL pathology.
Figure 2.Various metabolic mechanisms that can get altered to shape into FCHL phenotype.
FCHL and Metabolic Syndrome
Apart from establishing link between FCH and associated genetic and loci, there is also a clear evidence of overlapping features between metabolic syndrome (insulin resistance syndrome) and clinical and laboratory symptomology of FCHL. While some data contrast due to higher mortality for FCHL, still the two entities show co-occurrence, mixed traits and matching phenotype. 33 Santilli et al have identified the risk of cardio-metabolic disorders to become additive with both metabolic syndrome or FCHL traits only once these patients lost a so-called “protective gene cluster”, thus signifying that specific gene abnormalities are more associated with certain traits than simpler concept of monogenic inheritance. 34 The most associated USF1 on chromosome-1 is also linked with both metabolism of lipids and glucose like PPAR-gamma, HNA4a and PTEN, so it’s very probable that some of the common symptomatology of FCHL related to USF1 region with metabolic syndrome could be because of this multiple transcription factor regulation and associated signaling mechanisms. 35, Table 3 shows provides an insight into the biochemical alterations in FCHL in comparison to metabolic syndrome.
Provided the search on PubMed yielded some data as mentioned in Table 3, still we believe the evidence is not convincing to segregate the two phenotypically similar disorders. Only take home message here is the fact that the two disorders are overlapping with slightly higher gravity risk gravity attributed with underlying FCHL than the mere presence of metabolic syndrome symptomatology.
Table 3. Distinguishing metabolic syndrome from Familial Combined Hyperlipidemia (FCHL)| Ser | Contrast between FCHL and metabolic syndrome (MS) | Reference |
|---|---|---|
| 1 | Preadipocytes culture showed up-regulation of CD36 and FAT regardless of metabolic influences, indicating defects in adipose tissue metabolism | Meex et al (2005) 36 |
| 2. | TNF-alpha associated with FCHL only | Pauciullo et al (2007) 37 |
| 3. | sdLDLc specific for FCHL, and not for metabolic syndrome | Pauciullo et al (2009)38 |
| 4. | Vascular remodeling markers like TIMP-1, TIMP-2 and MMP-9 plasma levels were higher in metabolic syndrome than FCHL | Cicero et al (2007) 39 |
| 5. | PAI (Plasminogen activator inhibitor type-1) and MPO (Myeloperoxodase) activity are higher in FCHL than metabolic syndrome indicating its higher atherogenicity and atherothrombotic potential than MS. | Carratala et al (2018) 40 |
| 6. | Insulin resistance was found to be significantly higher in MS subjects with FCHL | Jackuliaková et al (2008) 41 |
Diagnosis of FCHL
While Framingham’s hyperlipidemia classification of type-2b with mixed hyperlipidemia is the initial clicker to include FCHL in your diagnosis, still we do learnt from above discussion that the FCHL phenotypes differ in terms of lipid levels, lipid types, associated metabolic component, clinical presentations, underlying genetics and inconsistent data on the subject. 5, 6 These considerations make the diagnosis of FCHL very much heterogeneous and a real life diagnostic dilemma for diagnosticians. Alongside we must appreciate the atherosclerotic cardiovascular disease associations of FCHL in terms of very high prevalence in patients having myocardial ischemia. 1 Sniderman et al have provided a general criteria for its diagnosis, but this still suffers with lack of specificity. 8 Not so recently we also had a nomogram for predicating susceptibility of FCHL as depicted in Figure 3, but unfortunately it could not gain widespread acceptability. 42
Figure 3.A nomogram suggested by Veerkamp et al to assess probability of having FCHL by incorporating data of Apolipoprotein-B, triglycerides and total cholesterol. Details for probability calculation are available at: https://www.ahajournals.org/doi/pdf/10.1161/01.CIR.0000130646.93255.86. Retrieved on: 19-Aug-2019.
Keeping in view the heterogeneity in FCHL phenotypes, the availability of various molecular tools and providing precise and accurate diagnosis it would be accurate to suggest that FCHL is convergence phenotype due to definitive molecular defects arising from a singular gene pathway like USF1 to genetic alterations across multiple other chromosomal regions as shown in Table 1. Furthermore, varying obesity patterns especially the “Asian obesity paradox” may not even present with hyperlipidemia, or sometimes as hypercholesterolemia or hypertriglyceridemia. 43
Molecular Diagnostics
Recent data in conventional medical science is being reshaped and supplanted with deeper insights into molecular physiology and pathological mechanisms. We are deemed to enter the arena of “molecular medicine”. We need to appreciate the fact that conventional medicine relies heavily on clinical phenotypes with biochemical evidence going alongside; however it can be said with confidence now that reliance on clinical phenotypes simply will not be able to provide personalized and nano-evidence based solutions for a specific patient. Further we move into the revolutionizing and innovative field of biotechnology, we are bound to find new tools and mechanistic set ups to better visualize the disease pathology. Below we attempt to provide a draft molecular diagnostic algorithm to approach subjects with either FCHL phenotype or a pattern like “Metabolic syndrome”. (Figure 4) This document is simply an attempt to combine new phenotype-genotype evidence to incorporate commonly appreciated molecular pathology on the subject. The limitations attached to this review must be understood: Firstly, the document is by no means intended to provide complete account of molecular defects due to evolving nature of both molecular pathology and discovery of newer dimensions of molecular pathology entering into the dimensions like miRNAs pathways, epigenetic mechanism and discovery of roles like “Alu element”. The data is bound to evolve and improve with further insight into complexities of various genetic pathways, miRNAs and related epigenetic methylation marks along with newer developments in nano-biotechnological tools. 44, 45 More so current text on metabolic syndrome and FCHL phenotype overlap is no more a mystery but we still need to appreciate the little differences in phenotypes with a specific genetic defect, for which the authors feel the genotype-phenotype data is limited and need more research.
Figure 4.Contributors including epigenetic, genetic, racial, regional factors leading to FCHL phenotype along with common molecular defects observed in metabolic syndrome.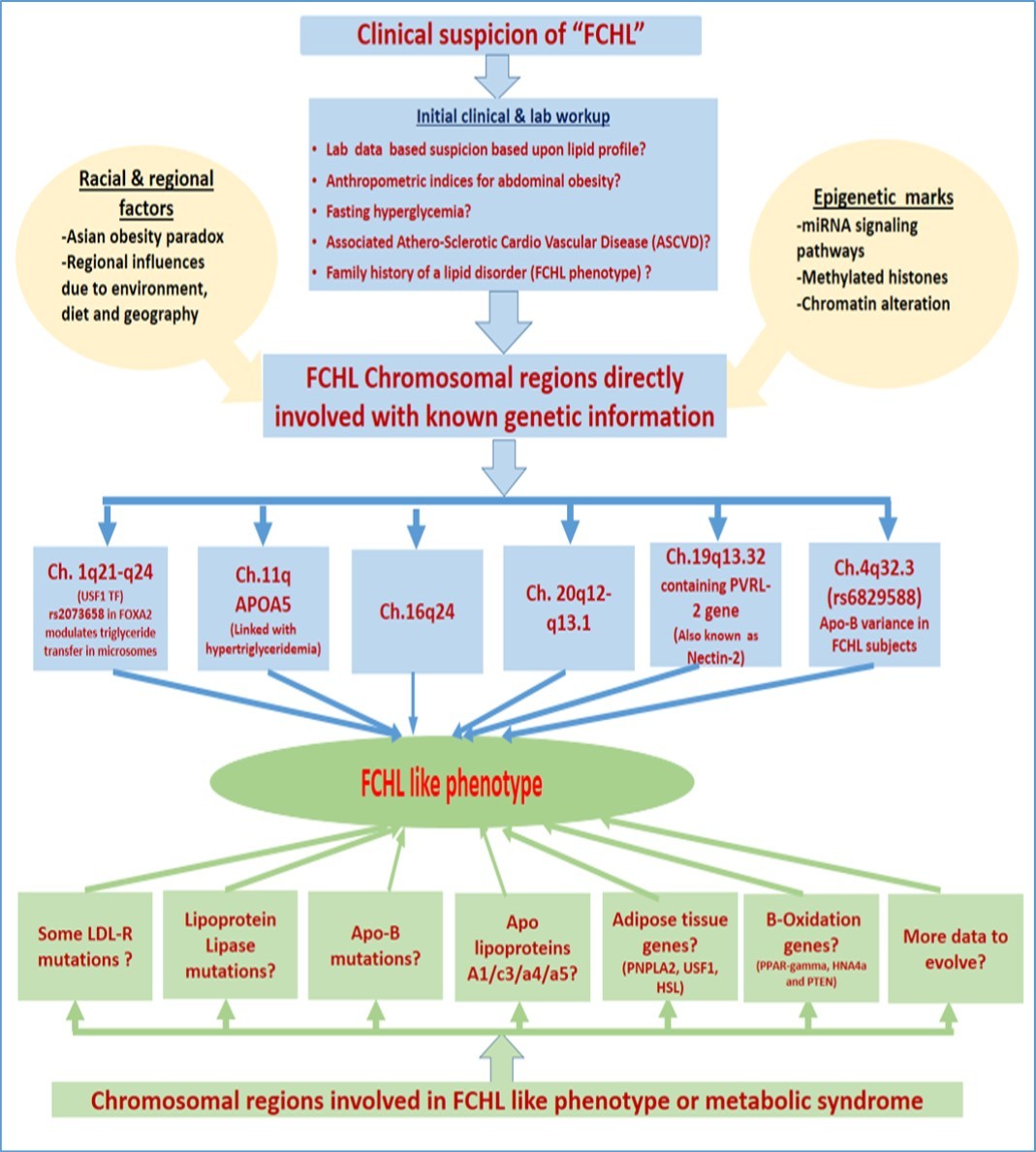
Provided certain limitations and lack of most recent molecular data on the subject, this review has clinical significance because it provided a wholesome overview of FCHL in terms of its contrasting phenotype molecular pathogenesis, polygenic nature of the disease, non-molecular metabolomics of FCHL within attempt to focus on futuristic way forward to diagnose and paving ways for gene therapy. It also emphasize the pushing needs for more research in defining further insights into FCHL with regards to emerging epigenetic markers like microRNAs, methylation studies and regional molecular differences within our community.
Conclusion
Molecular pathology is probably is the way to next generation medicine. The reviewed data suggested that FCHL phenotype is the resultant end outcome from multiple molecular defects and thus underlying genetic defect identification in the index case is important for personalized medicine and incoming gene therapy. Metabolic syndrome shares many of the symptoms and biochemical findings with FCHL phenotypes but still literature identifies differences. Our conclusion is that both metabolic syndrome and FCHL phenotype are broad group of disorders and molecular insight is needed to precisely and accurately identify the genetic or epigenetic defects in individuals for evidence based medicine and most importantly the gene therapy. We also feel this is an area of much needed research with detailed evaluation of genetic risk scoring for each genetic defect which may be possible in future.
References
- 1.Gaddi A, GalettiC PauciulloP, ArcaM. (1999) Familial combined hyperlipoproteinemia: expert’s panel position on diagnostic criteria for clinical practice. Committee of experts of the Atherosclerosis and Dysmetabolic Disorders Study Group.NutrMetabCardiovasc Dis.9(6): 304-11.
- 2.Brouwers M C, vanGreevenbroekMM StehouwerCD, J de Graaf, StalenhoefAF. (2012) The genetics of familial combined hyperlipidaemia.Nat. , Rev Endocrinol.Feb 8(6), 352-62.
- 3.WiesbauerF BlessbergerH, Azar D, Wagner O GoliaschG, GerholdL. (2009) Familial-combined hyperlipidaemia in very young myocardial infarction survivors (< or =40 years of age).Eur Heart. 30(9), 1073-9.
- 4.Brouwers M C, derKallenCJ Schaper NC van, vanGreevenbroekMM StehouwerCD. (2010) Five-year incidence of type 2 diabetes mellitus in patients with familial combined hyperlipidaemia. , Apr,NethJ 68(4), 163-7.
- 5.Ribalta J, La Ville AE, Heras M, Plana N, Masana L. (1997) Familial combined hyperlipidemia: detection and characterisation of the hyperlipidemic profile among children and adolescents. , Med Clin (Barc). Jun,28; 109(5), 161-4.
- 6.Bello-Chavolla O Y, Kuri-García A, Ríos-Ríos M, Vargas-Vázquez A, Cortés-Arroyo J E et al. (2018) . FAMILIAL COMBINED HYPERLIPIDEMIA: CURRENT KNOWLEDGE, PERSPECTIVES, AND CONTROVERSIES. Rev Invest Clin 70(5), 224-236.
- 7.Ellis K L, Pang J, Chan D C, Hooper A J, Bell D A et al. (2016) Familial combined hyperlipidemia and hyperlipoprotein(a) as phenotypic mimics of familial hypercholesterolemia: Frequencies, associations and predictions. J Clin Lipidol. 10(6), 1329-1337.
- 8.Sniderman A D, Castro Cabezas M, Ribalta J, Carmena R, de Bruin TW et al. (2002) A proposal to redefine familial combined hyperlipidaemia -- third workshop on. FCHL held in Barcelona from 3 to 5 , Eur J Clin Invest. Feb 32(2), 71-3.
- 9.Caron J, PèneV TolosaL, Luce E VillaretM, FourrierA. (2019) Low-density lipoprotein receptor-deficient hepatocytes differentiated from induced pluripotent stem cells allow familial hypercholesterolemia modeling, CRISPR/Cas-mediated genetic correction, and productive hepatitis C virus infection.Stem Cell ResTher.Jul,29;. 10(1), 221-10.
- 10.PajukantaP NuotioI, Terwilliger J D, Ylitalo K PorkkaKV, PihlajamäkiJ.Linkage of familial combined hyperlipidaemia to chromosome 1q21-q23.Nat. , Genet.1998 Apr; 18(4), 369-73.
- 11.Pei W, Baron H, Knoblauch H Müller-MyhsokB, Hui R Al-YahyaeeSA. (2000) Support for linkage of familial combined hyperlipidemia to chromosome. 1q21-q23 in Chinese and German families.Clin Genet.Jan; 57(1), 29-34.
- 12.vanGreevenbroekMM StalenhoefAF, J de Graaf, Brouwers M C. (2014) Familial combined hyperlipidemia: from molecular insights to tailored therapy.CurrOpinLipidol.Jun;. 25(3), 176-82.
- 14.J de Graaf, Vleuten G van der, StalenhoefAF. (2004) Diagnostic criteria in relation to the pathogenesis of familial combined. 4(3), 229-40.
- 15.Wijsman E M, Rothstein J H, Jr IgoRP, BrunzellJD MotulskyAG, JarvikGP. (2010) Linkage and association analyses identify a candidate region for apoB level on chromosome. 4q32.3 in FCHL families.Hum Genet.Jun; 127(6), 705-19.
- 16.SuviolahtiE LiljaHE, PajukantaP. (2006) Unraveling the complex genetics of familial combined hyperlipidemia.Ann Med.38(5):. 337-51.
- 17.Cruz-Bautista I, Mehta R, Cabiedes J, García-Ulloa C, Guillen-Pineda L E et al. (2015) Determinants of VLDL composition and apo B-containing particles in familial combined hyperlipidemia. Clin Chim Acta. 1, 160-165.
- 18.Minicocci I, Prisco C, Montali A, A Di Costanzo, Ceci F et al. (2015) Contribution of mutations in low density lipoprotein receptor (LDLR) and lipoprotein lipase (LPL) genes to familial combined hyperlipidemia (FCHL): a reappraisal by using a resequencing approach. , Atherosclerosis. Oct; 242(2), 618-24.
- 19.Brahm A J, Hegele R A. (2016) Combined hyperlipidemia: familial but not (usually) monogenic. , Curr Opin Lipidol. Apr; 27(2), 131-40.
- 20.Ellis K L, Pang J, Chan D C, Hooper A J, Bell D A et al. (2016) Familial combined hyperlipidemia and hyperlipoprotein(a) as phenotypic mimics of familial hypercholesterolemia: Frequencies, associations and predictions. , J Clin Lipidol. Nov - Dec; 10(6), 1329-1337.
- 21.Mo X, Lei S, Zhang Y, Zhang H. (2018) Genome-wide enrichment of m6A-associated single-nucleotide polymorphisms in the lipid loci. , Pharmacogenomics J. Sep 19(4), 347-357.
- 22.Auer S, Hahne P, Soyal S M, Felder T, Miller K et al. (2012) Potential role of upstream stimulatory factor 1 gene variant in familial combined hyperlipidemia and related disorders. Arterioscler Thromb Vasc Biol. 32(6), 1535-44.
- 23.Weissglas-Volkov D, Huertas-Vazquez A, Lee J SuviolahtiE, PlaisierC Canizales-QuinterosS. (2006) Common hepatic nuclear factor-4alpha variants are associated with high serum lipid levels and the metabolic syndrome.Diabetes.Jul;. 55(7), 1970-7.
- 24.TaghizadehE EsfehaniRJ, SahebkarA ParizadehSM, Rostami D, MirinezhadM. (2019) Familial combined hyperlipidemia: An overview of the underlying molecular mechanisms and therapeutic strategies.IUBMB Life.Sep;. 71(9), 1221-1229.
- 25.Díaz-Ruiz M, Martínez-Triguero M L, López-Ruiz A, F, Bañuls C et al. (2019) Metabolic disorders and inflammation are associated with familial combined hyperlipemia. , Clin Chim Acta. Mar; 490, 194-199.
- 26.Vlachopoulos C, Koutagiar I, Terentes-Printzios D, Skoumas I, Rigatou A et al. (2018) Relationship of PCSK9 levels with indices of vascular function and subclinical atherosclerosis in patients with familial dyslipidemias. Hellenic. 25, 1109-9666.
- 27.VaverkováH KarásekD. (2018) Familial combined hyperlipidemia - the most common genetic dyslipidemia in population and in patients with premature atherothrombotic cardiovascular disease.VnitrLek.Winter. 64(1), 25-29.
- 28.Carratala A, Martinez-Hervas S, Rodriguez-Borja E, Benito E, Real J T et al. (2018) PAI-1 levels are related to insulin resistance and carotid atherosclerosis in subjects with familial combined hyperlipidemia. , J Investig Med. Jan; 66(1), 17-21.
- 29.Arai H, Ishibashi.S,BujoH,Hayashi T,Yokoyama S,Oikawa S, et al; (2012).Research Committee for Primary Hyperlipidemia. Research on Measures against Intractable Diseases by the Ministry of Health,Labourand Welfare in Japan. Management of type IIb 19(2), 105-14.
- 30.Pei W D, Sun Y H, Lu B, Liu Q, Zhang C Y et al. (2007) Apolipoprotein B is associated with metabolic syndrome in Chinese families with familial combined hyperlipidemia, familial hypertriglyceridemia and familial hypercholesterolemia. , Int J Cardiol. Mar 116(2), 194-200.
- 31.AlipourA ElteJW, Rietveld AP vanZaanenHC, Cabezas M C. (2007) Postprandial inflammation and endothelial dysfuction.BiochemSoc Trans.Jun;. 35(3), 466-9.
- 32.Pei W D, Sun Y H, Rou W J, Zu Q, Li Y et al. (2005) Apolipoprotein B is associated with metabolic syndrome in Chinese pedigrees with familial hyperlipidemia. Zhonghua Yi Xue Za Zhi. , Feb 85(5), 313-7.
- 33.Cicero A F, Derosa G, Maffioli P, Reggi A, Grandi E et al. (2013) Influence of metabolic syndrome superposition on familial combined hyperlipoproteinemia cardiovascular complication rate. , Arch Med Sci. Apr 9(2), 238-42.
- 34.Santilli F, Blardi P, Scapellato C, Bocchia M, Guazzi G et al. (2015) Decreased plasma endogenous soluble RAGE, and enhanced adipokine secretion, oxidative stress and platelet/coagulative activation identify non-alcoholic fatty liver disease among patients with familial combined hyperlipidemia and/or metabolic syndrome. Vascul Pharmacol. 72, 16-24.
- 35.Rada-Iglesias A, AmeurA KapranovP, Enroth S, KomorowskiJ GingerasTR, WadeliusC. (2008) Whole-genome maps of USF1 and USF2 binding and histone H3 acetylation reveal new aspects of promoter structure and candidate genes for common human disorders.Genome Res.Mar;. 18(3), 380-92.
- 36.van derKallenCJ MeexSJ, Eurlings PM vanGreevenbroekMM, ElHasnaouiM EveloCT. (2005) Up-regulation of CD36/FAT. in preadipocytes in familial combined hyperlipidemia.FASEB 19(14), 2063-5.
- 37.Gentile M PauciulloP, Marotta G, BaianoA UbaldiS, JossaF. (2008) Tumor necrosis factor-alpha is a marker of familial combined hyperlipidemia, independently of metabolic syndrome.Metabolism.Apr;57(4):. 563-8.
- 38.Gentile M PauciulloP, Marotta G, BaianoA UbaldiS, JossaF. (2009) Small dense low-density lipoprotein in familial combined hyperlipidemia: Independent of metabolic syndrome and related to history of cardiovascular events.Atherosclerosis.Mar;203(1):. 320-4.
- 39.Cicero A F, Derosa G, Manca M, Bove M, Borghi C et al. (2007) Vascular remodeling and prothrombotic markers in subjects affected by familial combined hyperlipidemia and/or metabolic syndrome in primary prevention for cardiovascular disease. , Endothelium. Jul-Oct; 14(45), 193-8.
- 40.Carratala A, Martinez-Hervas S, Rodriguez-Borja E, Benito E, Real J T et al. (2018) PAI-1 levels are related to insulin resistance and carotid atherosclerosis in subjects with familial combined hyperlipidemia. , J Investig Med 66(1), 17-21.
- 41.JackuliakováD VaverkováH, KarásekD. (2008) Relationship between familial combined hyperlipidemia and insulin resistance.VnitrLek.Nov;54(11):. 1045-53.
- 42.VeerkampMJ de Graaf J, Hendriks J C, DemackerPN StalenhoefAF. (2004) Nomogram to diagnose familial combined hyperlipidemia on the basis of results of a 5-yearfollow-up study.Circulation. 109(24), 2980-5.
Cited by (2)
This article has been cited by 2 scholarly works according to:
Citing Articles:
V. Chernyshov - Ukrainian Therapeutical Journal (2021) Semantic Scholar
Ukrainian Therapeutical Journal (2021) OpenAlex
