Genetic Diversity, Phylogenetic Tree and Principal Component Analysis Based on Morpho-Metric Traits of Assam Chilli
Abstract
We evaluated a set of 37 chilli genotypes collected and maintained at Assam Agricultural University, Jorhat for 27 different traits related to plant habit (5), leaf (6), flower (2), fruit (13) and biotic stress (1). The variation in fruit yield among the genotypes could be attributed to high coefficients of variability for component traits viz., number of fruits per plant (91.7%), plant height (80.8%), leaf breadth (55.9 %), fruit weight (49.7%), leaf length (45.4%) fruit length (35.8%), fruit breadth (35.5%) and number of branches per plant (22.2%). Maximum phenotypic variants were observed for fruit traits followed by leaf characteristics. Phylogenetic analysis revealed Euclidean distances varying from a minimum of 2.065 and a maximum of 13.311 indicating the diverse nature of the genotypes. UPGMA clustering grouped the genotypes into 5 distinct clusters. The largest one, cluster I, had 26 genotypes belonging to Capsicum annuum var. acuminatum. Cluster II consisted of Capsicum annuum var. conoides with cone-shaped fruits. Cluster III included Moni Jolokia, a perennial shrub with cone-shaped globose erect fruits which clustered in between the other local C. annuum sp. Bireek and Mem Jolokia. The fourth cluster (IV) included the local chilli genotypes - Mem Jolokia, Bhekuri Jolokia and Haitha Jolokia which were perennial, with green stem and leaves. Cluster V included the C. chinense genotypes consisting of Manipuri Bhut, Bor Bhut and Lota Bhut. The first principal component explained 34.93% of the total variation contributed by mostly leaf and fruit characteristics. The fruit characters in this component showed significant positive correlation with leaf length, breadth and plant height indicating their importance in the morphological characterization of the chilli genotypes.
Author Contributions
Academic Editor: Mounira Elbaz, Regional Centre of Research on Horticulture and Organic Agriculture (CRRHAB) Chott Mariem, Sousse Tunisia · Horticulture, Tunisia
Checked for plagiarism: Yes
Review by: Single-blind
Copyright © 2018 Prabalee Sarmah, et al.
 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Competing interests
The authors have declared that no competing interests exist.
Citation:
Introduction
Chilli (Capsicum annuum L.) (2n = 24) of the Solanaceae family is a dicotyledonous flowering plant grown worldwide 1 and is a very important spice and vegetable crop valued for its aroma, taste, flavour and pungency. India is the leading chilli producing country in terms of area and production. The origin of this spice and vegetable crop can be traced to South and Central America 2. A few cultivars of hot pepper are cultivated outside their region of origin and nearly all those grown in tropics are local selections. C. annuum was domesticated in highlands of Mexico and includes Mexican, Asian and African hot chilli. There are taxonomic controversies regarding cultivated and wild forms of chillies but the most desirable forms are under C. annuum (both hot and sweet pepper) which are widely cultivated. Capsicum sp. are grouped under three species complexes- the C. annuum, C. baccatumand C. pubescence complexes. Among the five cultivated species of chilli, C. annuum, C. frutescence and C. chinenseJacq. belongs to the annuum complex.
Green fruits of chilli are used as vegetable whereas ripe dried fruits as a spice because of its pungency and pleasant flavour 3. Mature fruits of chilli contain carotenoid pigments, flavonoids and significant amounts of vitamin C, B1 and B2. The capsaicinoid compounds, which are responsible for the pungent flavour, are widely used for medicinal applications 4 such as cure of pharyngitis, lymph node swellings or boils and pain relief. In addition to the various medicinal uses, various forms of chilli with a whole host of colours also make it an important ornamental plant.
Plant biodiversity or the plant genetic wealth of an area is the variability between the plant kingdom and the ecosystem complex in which they occur. Capsicum is a highly polymorphic genus with much morphological and genetic diversity at both intra- and inter-specific level. The existence of extensive variability has been reported in local populations but with no such systematic report. In India, a rich diversity in chilli exists due to its diverse geo-climatic regions and the reproductive behaviour of the crop. Moreover, genetic diversity may arise due to the environmental influence and may be determined by the magnitude and nature of their genetic variability in which they are grown 5. Maximum diversity arises within cultivars grown in India with respect to shape, size, yield and other quality traits. This may be due to 2-40 % outcrossing which occurs due to entomophily. In order to access the source of genetic diversity adequate knowledge on the population structure of the particular species is very much essential. Diversity analysis among the working collections of germplasm enhances the efficacy of breeding and germplasm management. The knowledge on diversity and genetic distances of variable forms of the germplasm available at a particular region is a prerequisite for improving cultivars and developing cultivars suited for various needs. Development of improved high yielding varieties is based on only few parents and most of the useful genes present in local landraces and wild relatives will be lost. This will have undesirable consequences like genetic erosion and increased vulnerability to biotic and abiotic stresses. Conservation and sustainable use of the variable forms of chilli is the need of the hour for persistent improvement of this crop. Keeping in view the importance of diversity analysis, the present study was done to characterize the local chilli genotypes collected, selected and maintained at Assam Agricultural University, Jorhat.
Materials and Methods
Plant Materials
A set of thirty-seven Assam chilli genotypes was considered for morphological characterization which included 30 genotypes of Capsicum annuum, 3 belonging to Capsicum chinenseviz., Manipuri Bhut, Bor Bhut and Lota Bhut and 4 were unknown species. Among the thirty Capsicum annuum types K-L-1, K-L-2 , K-L-3, K-L-4, K-L-13 , K-L-14, K-L-31, K-L-35, K-S-4, K-S-9, K-S-11, K-S-32, K-S-37, K-S-38, K-S-41 ,K-S-47, Krishna L (different variable forms of black chilli), Khorika, KA-2, 2016/CHIVAR-2, CHIV AR-3, CHIVAR-4, CHIVAR-5, CHIVAR-6, CHIVAR-7 and CHIVAR-8 belonged to common chilli botanical variety acuminatum;Bireek-1, Bireek-2, B-1-P Sel, B1 x B2 belonged to cone chilli types Capsicum annuum var. conoides; and Bhekuri Jolokia to type cherry pepper Capsicum annuum var. cerasiforme. Two local chilli genotypes Mem Jolokia and Haitha Jolokia belongedto Capsicum annuum but with perennial growth habits. Except for 2016/CHIVAR-2, CHIVAR-3, CHIVAR-4, CHIVAR-5, CHIVAR-6, CHIVAR-7 and CHIVAR-8 which were provided by IIVR, Varanasi under AICRP (Vegetable Crops) varietal trial, and the other germplasm were locally collected from different places of Assam (Table 1).
Table 1. Details of the chilli genotypes with geographical coordinates| S No. | Genotype name | Botanical name | Pedigree | Origin | Geographical coordinates |
| 1-16 | K-L-1, K-L-2, K-L-3, K-L-4, K-L-13, K-L-14, K-L-31, K-L-35, K-S-4, K-S-9, K-S-11, K-S-11, K-S-32, K-S-37, K-S-38, K-S-41, K-S-47 | Capsicum annuum var acuminatum | Purelineselection from Krishna | Assam, India | 26 ̊ 47' Nlatitude94 ̊ 12' E longitude |
| 17 | Bireek-1 | Capsicum annuum var conoides | Landrace | Assam, India | 25 ̊ 82' N latitude93 ̊ 72' E longitude |
| 18 | Bireek-2 | Capsicum annuum var conoides | Landrace | Assam, India | Same as above |
| 19 | B-1-P Sel | Capsicum annuum var conoides | Purelineselection from Bireek 1 | - | 26 ̊ 47 'N latitude94 ̊ 12 ' E longitude |
| 20 | B1 x B2 | Capsicum annuum var conoides | F1 of Bireek 1/Bireek 2 | - | 26 ̊ 47 'N latitude 94 ̊ 12 ' E longitude |
| 21 | Krishna L | Capsicum annuum var acuminatum | Landrace | Assam, India | 26 ̊ 47 'N latitude 94 ̊ 10 ' E longitude |
| 22 | Khorika | Capsicum annuum var acuminatum | Landrace | Assam, India | 26 ̊ 47 ' N latitude94 ̊ 10 ' E longitude |
| 23 | KA-2 | Capsicum annuum var acuminatum | Unknown | - | - |
| 24 | Moni Jolokia | Capsicum sp. | Landrace | Assam, India | - |
| 25 | Manipuri Bhut | Capsicum chinense | Landrace | Manipur,India | 25 ̊ 12' N latitude93 ̊ 80' E longitude |
| 26 | Bor Bhut | Capsicum chinense | Landrace | Assam, India | - |
| 27 | Lota bhut | Capsicum chinense | Land race | Assam, India | - |
| 28 | Haitha Jolokia | Capsicum sp. | Landrace | Assam, India | 26 ̊ 47 'N latitude94 ̊ 13' E longitude |
| 29 | Bhekuri Jolokia | Capsicum sp. | Landrace | Assam, India | 26 ̊ 47 'N latitude 94 ̊ 13' E longitude |
| 30 | Mem Jolokia | Capsicum sp. | Land race | Assam, India | 26 ̊ 47'N latitude94 ̊ 12' E longitude |
| 31-37 | 2016/CHIVAR-2, 2016/CHIVAR-3, 2016/CHIVAR-4, 2016/CHIVAR-5, 2016/CHIVAR-6, 2016/CHIVAR-7, 2016/CHIVAR-8 | Capsicum annuum var acuminatum | Unknown | - | - |
Qualitative and Quantitative Observations
The experimental area is situated between 26o46/ N latitude and 94o13/ E longitude at an elevation of 86.56 m from the mean sea level. The soil of the experimental area belongs to Inceptisols derived from the alluvial deposits of the river Brahmaputra and its tributary Bhogdoi. The soil is light textured, acidic (pH: 4.8) and medium in organic matter content. Agricultural meteorological data during the experimentation period recorded an average maximum temperature of 27.22oC and a minimum temperature of 16.38oC, an average relative humidity of 94.6 percent and 20.98 mm rainfall.
The fertilizers were applied at 120:60:60 kg N:P2O5:K2O per ha in the form of Urea, Single Super Phosphate and Muriate of Potash at final land preparation. Irrigation was done as and when necessary. Prior to the experimentation, seeds of the genotypes were obtained from previous crops maintained through temporally and spatially isolated plots in poly houses.
Morphological data were obtained from the plants on the all thirty accessions assembled from different places of India. Twenty-seven different traits related to plant habit (5), leaf (6), flower (2), fruit (13) and biotic stress (1) were recorded as per NBPGR Minimal Descriptors for Agri-Horticultural Crops, Part-II: Vegetable Crops 6. Observations were taken on five randomly selected plants from the genotypes for the quantitative traits.
Statistical Analysis
The statistical calculations for mean, range, the coefficient of variation were performed in Microsoft Office Excel 2007. The qualitative and quantitative data were standardized using the mean and standard deviation of the samples and used to work out the Euclidian distances based dissimilarity matrix. An unrooted phylogenetic tree was then produced using Unweighted Paired Group Method using Arithmetic Average (UPGMA) of hierarchical clustering based on dissimilarity matrix as implemented in the DARwin software version 6.0. Principal component analysis (PCA) was performed following the method of 7 using the Analyse-it software Microsoft Excel 32 bit and the biplot diagram was generated using PC1 and PC2.
Results and Discussion
Mean Performance, Variability and Correlation
Abundant diversity was observed for all the quantitative traits considered in the study. Plant architecture was studied using plant height, number of branches per plant, leaf length and leaf breadth, plant growth habits, stem colour, leaf size, shape, margin and colour. Plant height ranged from 33 to 315 cm with a maximum height recorded in Manipuri Bhut (C. chinense) and a minimum plant height recorded in KA-2 (Capsicum annuum var. acuminatum). The number of primary branches varied from 3.6 to 11, with a maximum value being shown by Khorika (Capsicum annuum var. acuminatum) and a minimum by Lota bhut (C. chinense). Leaf length also showed a wide range from 3.6 to 17.5 cm. Maximum leaf length was observed in Manipuri Bhut (C. chinense) and the minimum leaf length was recorded in KS-38 (Capsicum annuum var. acuminatum). Similarly, leaf breadth also showed variation with values ranging from 1.4 to 7.6 cm. The broadest leaf was observed in Manipuri Bhut (C. chinense)while narrowest leaves were observed in Krishna L (Capsicum annuum var. acuminatum). Great diversity in fruit characters like fruit length, breadth, weight and number of fruits per plant was also observed. Fruit length, fruit breadth and fruit weight ranged from 1.32 to 10.2 cm, 0.37 to 2.23 cm and 1.5 to 10.2 g respectively. A wide diversity in the number of fruits per plant was observed ranging from 28 fruits in KL-2 to 846 fruits in Manipuri Bhut (C. chinense). Maximum yield per plant was obtained in Manipuri Bhut (C. chinense) 5761.26 g while the minimum yield of 146.4 g was observed in KS-11 (Capsicum annuum var. acuminatum). The maximum mean value was observed in yield per plant, followed by the number of fruits per plant and followed by plant height (Table 3). Based on the range and CV, the most variable character was fruit yield per plant (138.74%) followed by the number of fruits per plant (91.7%) and plant height (80.8%). The genotypes of C. chinense yielded higher than the C. annuum genotypes. The presence of wide range of variation in the component traits such as plant height (80.8%), number of branches per plant (22.2%), fruit length (35.8%), fruit breadth (35.5%), fruit weight (49.7%), number of fruits per plant (91.7%), leaf length (45.4 %) and leaf breadth (55.9 %), corroborated the greater yield differences among the genotypes.
Table 3. Range, mean and variation coefficient for the nine quantitative characters| Character | Range | Mean ± SEm | CV (%) |
| Plant height (cm) | 33.00 - 315.40 | 80.71 ± 10.87 | 80.77 |
| Number of branches/plant | 3.60 - 11.00 | 6.63 ± 0.25 | 22.22 |
| Leaf length (cm) | 3.60 - 17.50 | 7.16 ± 0.54 | 45.38 |
| Leaf breadth (cm) | 1.42 - 7.60 | 2.97 ± 0.28 | 55.9 |
| Fruit length (cm) | 1.32 - 10.20 | 6.14 ± 0.37 | 35.84 |
| Fruit breadth (cm) | 0.37 - 2.23 | 1.27 ± 0.07 | 35.48 |
| Fruit weight (g) | 1.50 - 10.20 | 4.38 ± 0.36 | 49.74 |
| Number of fruits/plant | 28.00 -846.00 | 184.94 ± 31.59 | 91.66 |
| Fruit yield/plant (g) | 146.40 -5761.26 | 762.77 ± 172.28 | 138.74 |
Pearson’s correlation coefficient among the quantitative traits revealed a significant positive correlation of plant height (PH), leaf length (LL), leaf breadth (LB) and number of fruits per plant (FN) with fruit yield in the chilli genotypes (Table 4). This suggested that the genotypes which were tall, with longer and broader leaves bore more number of fruits giving higher yield. These traits were also important for distinguishing the genotypes in the species and sub-species level. Significant negative correlation between fruit breadth and number of branches per plant indicated that the genotypes with more number of branches bore fruits with narrower fruit breadth. These traits could be important criteria for improvement of yield in chilli.
Table 4. Pearson correlation coefficients among the quantitative traits in chilli| Character | PH | BR | LL | LB | FL | FB | FW | FN |
| BR | -0.47 | |||||||
| LL | 0.87 | -0.49 | ||||||
| LB | 0.73 | -0.43 | 0.91 | |||||
| FL | -0.47 | 0.52 | -0.42 | -0.49 | ||||
| FB | 0.52 | -0.56 | 0.47 | 0.38 | -0.14 | |||
| FW | -0.01 | -0.02 | 0.12 | -0.01 | 0.46 | 0.36 | ||
| FN | 0.71 | -0.20 | 0.73 | 0.74 | -0.48 | 0.20 | -0.11 | |
| FY | 0.79 | -0.24 | 0.77 | 0.62 | -0.20 | 0.41 | 0.29 | 0.81 |
| PH: Plant height (cm); BR: Branches per plant; LL: Leaf length (cm); LB: Leaf breadth (cm); FL: Fruit length (cm); FB: Fruit breadth (cm); FW: Average fruit weight (g); FN: Fruit number per plant; FY: Fruit yield per plant (g) | ||||||||
The number of phenotypic variants for the 37 genotypes as per NBPGR Minimal Descriptors for Horticultural Crops revealed a lot of variation for the studied traits (Table 2). Among the eighteen qualitative characters, an annual life cycle was exhibited by all the genotypes except the C. chinense genotypes, Moni Jolokia, Bhekuri Jolokia, Mem Jolokia and Haitha Jolokia which were semi-perennial to perennial. Stem colour showed 17 genotypes with green phenotype, 11 with purple and 9 with green with purple stripes. Plant growth habit among the 37 genotypes exhibited 17 with prostate, 13 with erect and 7 with intermediate growth.
Table 2. Phenotypic variants observed for the various qualitative traits across the 37 genotypes| S No. | Trait | Score | Phenotype | No. of genotypes | S No. | Trait | Score | Phenotype | No. of genotypes |
| 1 | Life cycle | 1 | Annual | 32 | 11 | Ripe fruit colour | 3 | Pale Orange Yellow | - |
| 2 | Biennial | - | 4 | Orange Yellow | 1 | ||||
| 3 | Perennial | 5 | 5 | Pale Orange | 2 | ||||
| 2 | Stem colour | 1 | Green | 17 | 6 | Orange | - | ||
| 2 | Green with purple stripes | 9 | 7 | Light Red | 4 | ||||
| 3 | Purple | 11 | 8 | Red | 8 | ||||
| 3 | Plant growth habit | 3 | Prostrate | 17 | 9 | Dark Red | 19 | ||
| 5 | Intermediate | 7 | 10 | Brown | 2 | ||||
| 7 | Erect | 13 | 11 | Purple | - | ||||
| 4 | Leaf size | 3 | Small | 16 | 12 | Black | - | ||
| 5 | Medium | 15 | 12 | Fruit shape | 1 | Long | 27 | ||
| 7 | Large | 6 | 2 | Very short | 2 | ||||
| 5 | Leaf shape | 1 | Deltoid | - | 3 | Tapering | 1 | ||
| 2 | Ovate | 13 | 4 | Conical | 3 | ||||
| 3 | Lanceolate | 24 | 5 | Oval | 4 | ||||
| 6 | Leaf margin | 1 | Entire | 35 | 13 | Fruit shape at pedicel attachment | 1 | Acute | 10 |
| 2 | Undulate | 2 | 2 | Obtuse | 18 | ||||
| 3 | Ciliate | - | 3 | Truncate | 8 | ||||
| 7 | Leaf colour | 1 | Green | 21 | 4 | Cordate | 1 | ||
| 2 | Dark Green | 10 | 5 | Lobate | - | ||||
| 3 | Purple | 6 | 14 | Fruit shape at the blossom end | 1 | Pointed | 24 | ||
| 8 | Corolla colour | 1 | White | 25 | 2 | Blunt | 9 | ||
| 2 | Yellow | - | 3 | Shrunken | - | ||||
| 3 | Purple | 12 | 4 | Shrunken and pointed | 4 | ||||
| 9 | Anther colour | 1 | White | 1 | 15 | Fruit position | 3 | Pendant | 23 |
| 2 | Yellow | 1 | 5 | Semi Pendent | 3 | ||||
| 3 | Pale Blue | 14 | 7 | Erect | 11 | ||||
| 4 | Blue | 4 | 16 | Adherence of calyx to fruit | 3 | Loose | 11 | ||
| 5 | Bluish Yellow | 1 | 5 | Semi-hard | 24 | ||||
| 6 | Purple | 16 | 7 | Hard | 2 | ||||
| 10 | Mature fruit colour | 1 | White | 1 | 17 | Fruit surface | 1 | Smooth | 19 |
| 2 | Yellow | - | 2 | Semi wrinkled | 15 | ||||
| 3 | Green | 15 | 3 | Wrinkled | 3 | ||||
| 4 | Orange | - | 18 | Biotic stress | 1 | Very low | 26 | ||
| 5 | Purple | 8 | 3 | Low | 10 | ||||
| 6 | Deep Purple | 1 | 5 | Intermediate | 1 | ||||
| 7 | Black | 12 | 7 | High | - | ||||
| 11 | Ripe fruit colour | 1 | White | - | 9 | Very high | - | ||
| 2 | Lemon Yellow | 1 | |||||||
Among the leaf characteristics, the maximum number of phenotypic variants were found for leaf size (16 small, 15 medium and 6 large) and colour (21 green, 10 dark green and 6 purple). Phenotypic variation for leaf shape gave 24 variants with lanceolate leaves and 13 with ovate leaves and 35 variants had entire leaf margin and 2 had undulating leaf margin. Similarly, for flower characteristics, 25 phenotypic variants were observed with white corolla and 12 variants were with different shades of a purple corolla. A wide variation was observed for anther colour showing phenotypic variants with 1 white, 1 yellow, 14 pale blue, 4 blue, 1 bluish yellow and 16 purple anther.
Among fruit characteristics, fruit colour exhibited enormous variation among genotypes. Phenotypic variants for fruit colour were 1 white, 15 green, 8 purple, 1 deep purple, 12 black matured, 1 lemon yellow, 1 orange-yellow, 2 pale orange, 4 light red, 8 red, 19 dark red and 2 brown ripe. Fruit shape variants among genotypes included 27 long, 2 very short, 1 tapering, 3 conical and 4 oval. The number of genotypes with variation in fruit shape at pedicel attachment were 10 acute, 18 obtuse, 8 truncate and 1 cordate. The blossom end fruit shape was observed to be pointed in 24, blunt in 9 and shrunken-pointed in 4 genotypes. Twenty-three of the phenotypic variants held a pendant position in the stem followed by erect and semi-pendent in 11 and 3 genotypes, respectively. The adherence of calyx to fruits was loose for 11, semi-hard for 24 and hard for 2 genotypes. The fruit surface was found to be smooth, semi-wrinkled and wrinkled for 19, 15 and 3 genotypes, respectively. A very low to intermediate biotic stress was observed in the genotypes with very low to no incidence in 26 variants, low incidence in 10 and intermediate incidence in 1 variant.
Phylogenetic Analysis
Phylogenetic analysis was performed for the 37 genotypes using standardized values for the qualitative and quantitative traits. Dissimilarity matrix of usual Euclidean distances gave a minimum dissimilarity value of 2.065 and a maximum of 13.311 indicating the diverse nature of genotypes under study. The unrooted phylogenetic tree constructed with dissimilarity values using UPGMA method grouped the genotypes into five major clusters (I, II, III, IV and V) clockwise from right to left (Figure 1). A big clustering (cluster I) was formed at node no 67 with the genotypes belonging to Capsicum annuum var. acuminatum. Based on the nodal separation of the branches of this cluster (I) at nodes 64, 65 and 66, it was further separated into three sub-clusters. The sub-cluster formed at node 66 comprised of Krishna L and eight other genotypes all having mature fruit colour purple to black and other similar fruit characteristics, similar plant architecture and leaf characteristics. It is remarkable to note that this cluster comprised of the genotypes with anthocyanin pigmentation in their stems and leaves to varying degree (Plate 1). Flowers showed different shades of purple. Fruits borne were pendent and long with pointed tips (Plate 2). These variant forms might have arisen due to natural outcrossing with Krishna L which is a landrace of Assam, India. This landrace has been under cultivation from ages and is a heavy bearer of pungent dark purple to black coloured matured fruits. The sub-cluster at node 64 comprised of the variant types of purple to black chilli genotypes, with long, semi-wrinkled pointed tip fruits, borne erect to semi-pendent on the stems. The flowers were with white corolla with a purple tinge and purple anthers. Green chilli genotypes formed sub-cluster at node 65. The genotypes bore fruits which were long, tapering with pointed ends and acute shape at pedicel attachment (Plate 2). These genotypes were green with very less purple pigmentation. Leaves were mostly lanceolate and smaller in size. Flowers were with white corolla and pale blue to purple anthers.
Figure 1.Unrooted phylogenetic tree of chilli genotypes based on qualitative and quantitative traits (The tree was generated using UPGMA based on the dissimilarity among the 37 chilli genotypes)
Cluster II at node no. 69 in the unrooted phylogenetic tree (Figure 1) consisted of the genotypes of Capsicum annuum var. conoides collected from Karbi Anglong district of Assam, India and two genotypes developed at AAU, Jorhat, Assam, India. Fruits were cone-shaped, borne erect on the pedicel, both green and purple in colour (Plate 1 and plate 2). These genotypes were tall with a lesser number of branches per plant; green stems and leaves. Cluster III (node no. 71) included one genotype, Moni Jolokia which was a perennial shrub, bearing cone-shaped globose erect fruits (Plate 1 and plate 2). The genotype bore big ovate leaves which were green in colour. The corollas were white with greyish blue anthers. This genotype was found to be clustering in between the other local C. annuum sp. Bireek and Mem Jolokia, but was not seen to form groups with these genotypes. The fourth cluster (IV) at node no. 72 included the local chilli genotypes Mem Jolokia, Bhekuri Jolokia and Haitha Jolokia. These three genotypes were perennial, with green stem and leaves. The two genotypes Mem Jolokia and Bhekuri Jolokia bore erect fruits, while Haitha Jolokia bore semi-pendent fruits (Plate 1). Cluster V included the C. chinense genotypes consisting of Manipuri Bhut, Bor Bhut and Lota Bhut (Plate 1 and plate 2). The presence of maximum diversity in the genotypes belonging to C. annuum was in accordance with the findings of 8.
Plate 1.Plant types of genotypes in clusters I(1-14), II(15), III(17), IV(16,20) AND V(18,19)
Plate 2.Fruit and leaf type of some genotypes in Cluster I
Plate 3.Flower, fruit and leaf types of some genotypes in different clusters
The plant architecture, leaf, flowers and fruit characteristic distinctly separated the genotypes belonging to the species Capsicum chinense from the other capsicum genotypes. The observed quantitative traits were also variable from the other capsicum genotypes. These genotypes showed a perennial growth habit and were high yielder bearing distinct, tapering, wrinkled surface, orange coloured ripe, highly pungent fruits (Plate 3). The UPGMA cluster analysis assumes a constant evolutionary rate among accessions and is typically most appropriate for diversity study within species 9. The clustering of the genotypes showed that within C. annuum genotypes, three sub-clusters originated based on the minor morphological variations. It was remarkable to note that Capsicum annuum var. acuminatum and Capsicum annuum var. conoides were found to be closely related to each other based on stem, leaf, flower and fruit characteristic (Plate 2) except that the genotypes under var.conoideswhich were late maturing as compared to the genotypes of var. acuminatum. Moreover, fruit surface, position and shape of the fruits of the var. conoides were quite distinct from those of the var. acuminatum (Plate 1 and plate 2). The local chilli genotypes Moni, Mem, Bhekuri and Haitha appeared to be close to the C. annuum genotypes and might be used as a source for gene introgression into the cultivated types as these local genotypes are rich sources of many useful genes including resistance to biotic and abiotic stresses as revealed by resistant performances during stresses both ex situ and in situ. These species also appeared to be in between the C. annuum and C. chinense genotypes which could be a potential source for chilli improvement as a whole. Among the domesticated species of Capsicum, C. chinensehas been reported to have a better crossability with C. annuum 10, 11, providing scope for chilli improvement. The genotypes of these two species could be used as diverse parents for the transfer of many desirable traits. Similarly, the two landraces Khorika and Krishna L also appeared to be dissimilar for many morphological traits. So even when placed in one big cluster I (at node 67), they were grouped in two separate sub-clusters. These genotypes exhibited good adaptability with respect to biotic and abiotic stresses and were high yielders. Thus, the cluster analysis resulted in a vivid picture implicating the usefulness of the genotypes for crop improvement programme. In an analogous study on the phenotypic divergence among 24 accessions belonging to six species of Capsicum, using 12 quantitative and qualitative traits, resulting data grouped the species into six clusters. Three traits among these were the most discriminative of these species, i.e., fruit diameter, number of fruits per plant and leaf diameter. They were reported useful as morphological markers for yield improvement and for obtaining good segregants in chilli breeding programs 12.
Principal Component Analysis (PCA)
PCA explained the proportion of relative contribution of the various characters to the total variance of chilli genotypes under study. The study of many qualitative and quantitative traits is important for the assessment of differences between genotypes and their breeding potential. PCA combines the capacity to provide a synthetic summary of the most relevant traits and assessment of the relative contribution of different characters to the total variability of the population 13. The PCA contributed 99.7 % to the total variability of chilli genotypes up to twenty-two principal components. The largest contribution to variability (34.93%) was made by the first component and smallest contribution (1%) by the twenty-second component. The first six components accounted for 81.44% of the total variability and these six components were considered for discussion.
The highest proportion of total variability exposed by the first component (34.93%) was contributed by a large number of traits with high positive values - plant height, leaf length, leaf breadth, fruit number per plant, fruit yield, fruit shape at blossom end, leaf size, life cycle and traits with low positive values - leaf margin, fruit shape, fruit shape at pedicel attachment and fruit surface (Table 5). The variability in this component was mostly linked to leaf and fruit characteristics which were mostly quantitative in nature. Correlation matrix of these traits revealed that fruit characters in this component showed significant positive correlation with leaf length, breadth and plant height suggesting that these characteristics could complement each other, and also appeared to be important for morphological characterization of the chilli genotypes. The high negative values in this component were observed for mature fruit colour, fruit length and branches per plant.
Table 5. Matrix of eigenvectors and values of principal components of the 27 chilli traits| Trait | Principal component | |||||
| C1 | C2 | C3 | C4 | C5 | C6 | |
| Plant height (cm) | 0.275 | 0.166 | -0.002 | 0.153 | -0.024 | 0.17 |
| No of branches per plant | -0.193 | -0.038 | 0.33 | 0.149 | -0.002 | -0.004 |
| Leaf length (cm) | 0.294 | 0.135 | 0.01 | 0.103 | 0.11 | -0.113 |
| Leaf breadth (cm) | 0.289 | 0.013 | 0.02 | 0.045 | 0.28 | -0.136 |
| Fruit length (cm) | -0.211 | 0.19 | 0.238 | -0.032 | 0.122 | -0.182 |
| Fruit breadth (cm) | 0.14 | 0.343 | -0.138 | -0.12 | 0.048 | -0.01 |
| Fruit weight (g) | -0.008 | 0.342 | 0.119 | -0.257 | 0.153 | -0.235 |
| No of fruits per plant | 0.266 | 0 | 0.108 | 0.205 | 0.035 | 0.125 |
| Fruit yield per plant (g) | 0.252 | 0.196 | 0.138 | 0.11 | -0.041 | 0.082 |
| Lifecycle | 0.286 | -0.098 | -0.001 | 0.059 | 0.205 | 0.096 |
| Stem colour | -0.174 | 0.18 | -0.261 | 0.079 | 0.209 | 0.04 |
| Plant growth habit | -0.047 | -0.191 | -0.348 | 0.223 | -0.087 | 0.246 |
| Leaf size | 0.258 | -0.072 | -0.103 | 0.053 | 0.13 | -0.257 |
| Leaf shape | -0.095 | 0.202 | 0.252 | 0.252 | -0.112 | 0.294 |
| Leaf margin | 0.102 | -0.302 | -0.057 | 0.126 | 0.427 | -0.038 |
| Leaf colour | -0.036 | 0.192 | -0.209 | 0.368 | 0.181 | 0.091 |
| Corolla colour | -0.154 | 0.161 | -0.247 | 0.28 | 0.138 | 0.111 |
| Anther colour | -0.176 | 0.227 | -0.266 | 0.006 | 0.24 | -0.06 |
| Mature fruit colour | -0.212 | 0.259 | -0.085 | -0.073 | 0.14 | -0.085 |
| Ripe fruit colour | -0.17 | 0.181 | -0.21 | -0.182 | -0.067 | 0.123 |
| Fruit shape | 0.167 | 0.091 | -0.332 | -0.071 | -0.357 | -0.075 |
| Fruit shape at pedicel attachment | 0.161 | 0.123 | -0.092 | -0.419 | 0.114 | 0.292 |
| Fruit shape at blossom end | 0.278 | 0.14 | -0.149 | -0.014 | -0.057 | 0.053 |
| Fruit position | 0.044 | -0.272 | -0.267 | -0.246 | -0.006 | -0.314 |
| Adherence of calyx to fruit | -0.099 | -0.159 | 0.113 | -0.211 | 0.532 | 0.274 |
| Fruit surface | 0.155 | 0.262 | 0.206 | 0.059 | 0.013 | -0.274 |
| Biotic stress | 0.146 | 0.048 | 0.096 | -0.35 | -0.022 | 0.463 |
| Total variance | 9.43 | 4.052 | 3.446 | 2.17 | 1.643 | 1.247 |
| Variance explained (%) | 34.926 | 15.008 | 12.763 | 8.039 | 6.085 | 4.619 |
| Cumulative (%) | 34.926 | 49.934 | 62.697 | 70.736 | 76.821 | 81.44 |
About 15% of the variability was contributed by the second principal component with high positive values for fruit breadth and fruit weight, and low positive values for plant height, leaf length, fruit length, fruit yield, stem colour, leaf shape, leaf colour, corolla colour, colour of matured fruit/ripe fruits, fruit surface, fruit shape at pedicel attachment and at blossom end, and fruit shape in general (Table 5). High negative values were obtained for leaf margin and fruit position. The correlation matrix (Table 4) also showed a significant positive correlation of plant height with leaf length, leaf breadth, fruit breadth, fruit yield per plant, fruit shape at the blossom end and fruit surface, suggesting effective complementation among the traits useful for crop improvement. The third PC contributing 12.76 % to the total variability showed a greater influence of the plant traits viz., number of branches per plant, leaf shape and fruit surface. The fourth and the fifth PC contributed 8.04% (leaf shape, leaf colour, corolla colour, number of fruits per plant, plant growth habit) and 6.08 % (leaf breadth, leaf margin, adherence of calyx to fruits, anther colour, stem colour), respectively to the variability (Table 5). The variability of these components was mostly linked to the qualitative traits and a few quantitative traits. Morphological characterization of 18 C. annuum L. accessions from southern Mexico using 47 morphological descriptors and response to Bemisiatabaci - Begomovirus complex resulted in 12 principal components accounting for 94% of the variation with 3 major clusters and 7 sub-clusters 14.
The PCA biplot diagram of the species drawn on the basis of PC1 and PC2, accounting for a total of 49.9% total variability (Figure 2) identified the traits that contributed profoundly to PC1 (plant height 83%, fruit shape at blossom end 81%, leaf length and life cycle 80%, leaf breadth 79%, colour of matured fruits 69% and fruit number 67%) in characterizing the different genotypes. These traits contributing to the PC1 (plotted on X-axis) distinctly grouped all the genotypes belonging to Capsicum annuum in a wide grouping. The genotypes of Capsicum annuum var. acuminatum and Capsicum annuum var. conoides did not show separate grouping. The local chilli genotypes Bhekuri, Haitha and Mem Jolokia were plotted near the Capsicum annuum group but at a distance. PCA biplot diagram plotted Mem Jolokia near the Capsicum annuum var. acuminatum genotypes as it might be an ecotype-specific genotype which was tall and perennial bearing innumerable, small and erect fruits. Haitha and Bhekuri were plotted at a distance which appeared to be the ecological variants with similar leaf patterns, but with vast differences in the stem, flower and fruit characteristics than Capsicum annuum. Haitha bore characteristic of semi-pendent flowers with white corolla, light green matured fruits and lemon yellow-ripe fruits whereas, Bhekuri was characterised by purple pigmentation in the stem, small white erect flowers and unique small, erect, globose fruits. The colour transformation in the fruits of this type was remarkable as immature fruits bore purplish tinge which gradually turned to white, matured fruits turned to yellow and finally turned red on ripening. The PCA biplot isolated the two C. chinense genotypes, Manipuri bhut and Bor bhut into a separate group perhaps because of the similarities in most of the morphological traits of these two genotypes except for some minor variations. Moni Jolokia was unique from the other groups but was close to the C. chinense genotypes most probably because of the similarities in fruit characteristics like fruits per plant, leaf characteristics and life cycle. This grouping was in accordance with cluster analysis with one exception C. chinense genotype Lota bhut which was separated from its group and was placed quite near a C. annuum genotype KA-2. These traits appeared to be important in characterizing the genotypes at the species level. The maximum contribution to PC2 was made by fruit yield (76%) followed by fruit breadth (66%) (Figure 2 ). The various traits explained by PC1 and PC2 was capable of resolving the genotypes to their respective species level. Cluster analysis and principal component analysis were used to group 54 chilli genotypes (Capsicum annuum L.) into seven clusters 3. Based on mean, variance and group distance, in his study, clusters I and III along with other clusters were suggested for hybridization programme. In the present study, the grouping of the chilli genotypes based on morphological characteristics to their respective species level through PCA analysis and cluster analysis was in general agreement.
Figure 2.PCA Biplot diagram of chilli genotypes based on PC 1 and PC 2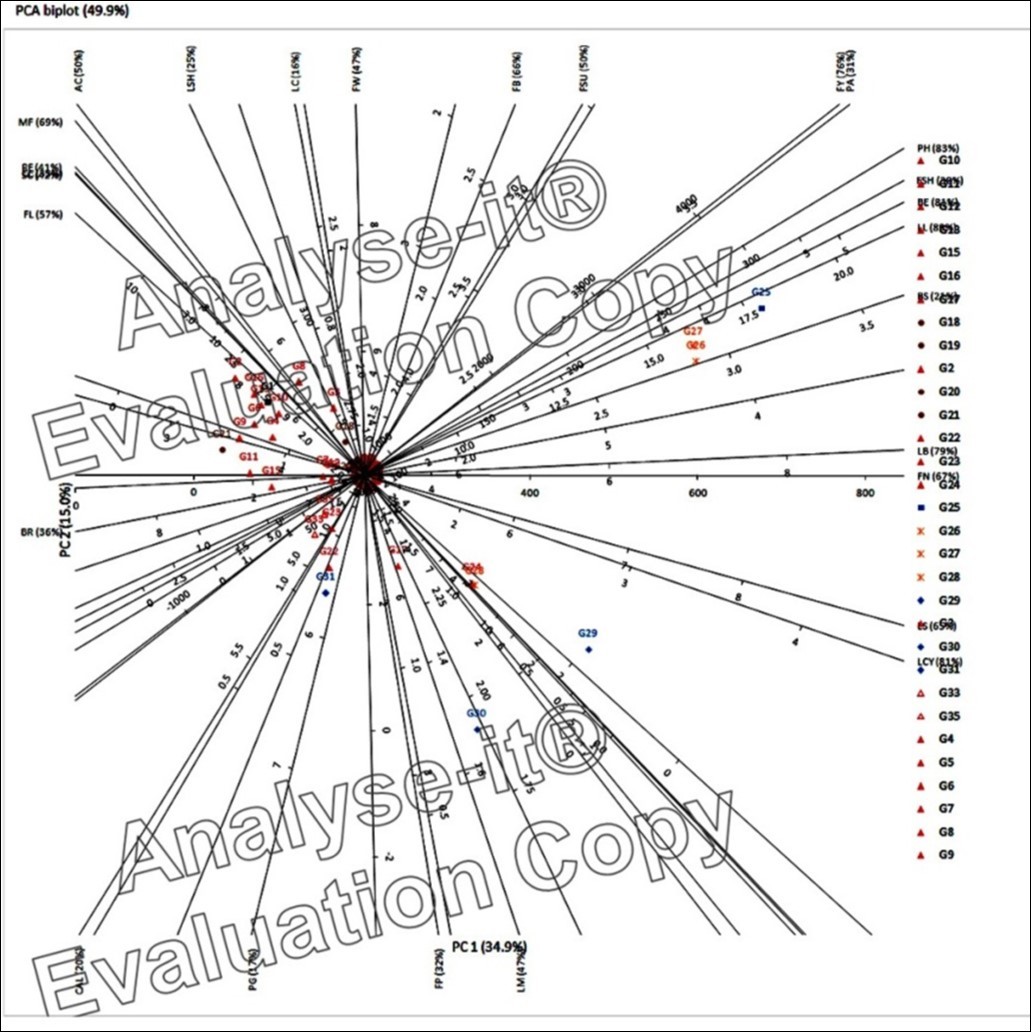
Conclusion
The results presented here demonstrate the utility of cluster analysis and PCA in partitioning the genetic variation among chilli genotypes and in identifying different genotypes of chilli which would serve as potential sources of unique breeding material for future crop improvement. The study also provides guidance for future analysis of genetic diversity using the more reliable molecular markers to facilitate efficient management and utilization of the available germplasm. Genetic fingerprinting of local germplasm would protect the plant genetic resources of the nation in light of the PPVFRA and Convention on Biological Diversity. A core collection could also be created with the morphological diversity for utilization in the future breeding programme.
Acknowledgement
The first author thankfully acknowledges the help provided by All India Coordinated Research Project on Vegetable Crops (ICAR), IIVR, Varanasi, India for providing some of the materials for the study and also for financial assistance.
References
- 1.Knapp S, Bohs L, Nee M, Spooner D M. (2004) Solanaceae- a model for linking genomics with biodiversity. , Comparative and Functional Genomics 5, 285-291.
- 2.Bahurupe J V, Sakhare S B, Kulwal P L, Akhare A A, Pawar B D. (2013) Genetic diversity analysis in chilli (Capsicum annuumL.) using RAPD markers. , The Bioscan 8(3), 915-918.
- 3.Hasan M J, Kulsum M U, Uttah M Z, Hossain M M, Mahmud M E. (2014) Genetic diversity of some chilli (Capsicum annuumL.) genotypes. , Int. J. Agril. Res. Innov. & Tech 4(1), 32-35.
- 4.Boslan P W, Votova E J. (2000) Peppers: Vegetable and spice capsicums. 2ndEd. Wallingford: Oxon, U.K.New York.Cabi-Crop Production Science in Horticulture Series:v2.(CrossRef) .
- 5.Falconer D S, TFC Mackay. (1996) Introduction to Quantitative Genetics. 4thEd.,Adison Wesley Longman,Harlow.
- 6.Srivastava U, Mahajan R K, Gangopadhyay K K, Singh M, Dhillon B S. (2001) Minimal Descriptors for Agri-Horticultural Crops, Published by. National Agricultural Technology Project on Plant Biodiversity (NATP-PB) and National Bureau of Plant Genetic Resources, Pusa Campus , New Delhi 39-45.
- 8.Lee Hea-Young, Venkatesh Jelli, Jung Ayoung, Han Ji-Woong, Yeaseong H A et al. (2016) Genetic diversity and population structure analysis to construct a core collection from a large Capsicum germplasm. , Doi: 10.1186/s12863-016-0452-8. BMC Genet 17, 142.
- 9.Sarmah P, Barua P K, Sarma R N, Sen P, Deka P C. (2007) Genetic diversity among rattan genotypes from India based on RAPD-marker analysis. , Genet Resour Crop Evol 54, 593-600.
- 10.Manzur J P, Fita A, Prohens J, Rodríguez-Burruezo A. (2015) Successful wide hybridization and introgression breeding in a diverse set of common peppers (Capsicum annuum) using different cultivated Ají (C.baccatum) accessions as donor parents.Doi: 10.1371/journal.pone.0144142. PLoS ONE 10:e0144142. (CrossRef)
- 11.Martins K C, Nair T, Pereira S, Alessandro S, Souza M et al. (2015) Crossability and evaluation of incompatibility barriers in crosses betweenCapsicumspecies. , Crop Breed Appl Biotechnol.(CrossRef) 15, 139-145.
- 12.Thul S T, Lal R K, Shasany A K, Darokar M P, Gupta A K et al. (2009) Estimation of phenotypic divergence in a collection of capsicum species for yield related traits. , Euphytica 168(2), 189-196.
Cited by (9)
This article has been cited by 9 scholarly works according to:
Citing Articles:
PeerJ (2023) OpenAlex
Sushil Kumar, J. Baruah, S. Munda, Neelav Sarma, Twahira Begum et al. - PeerJ (2023) Semantic Scholar
PeerJ (2023) Crossref
Acta Horticulturae (2023) OpenAlex
V.K. Sharma, A. Srivastava, M. Mangal, C.D. Pandey, V.P. Sharma et al. - Acta Horticulturae (2023) Semantic Scholar
Acta Horticulturae (2023) Crossref
Sarhad Journal of Agriculture (2022) OpenAlex
P. Rahevar, J. N. Patel, Axatjoshi, Sushilkumar, Lalji N. Gediya - (2021) Semantic Scholar
Electronic Journal of Plant Breeding (2021) OpenAlex
