Polysaccharide Transglycosylases: A Survey of Assay Methods
Abstract
Polysaccharide transglycosylases (PTGs) are a unique group of glycoside hydrolases playing important roles in the formation and modification of plant and fungal cell walls. Their action involves cutting the molecule of the polysaccharide substrate at the glycosidic bond, followed by transfer of the newly formed reducing-end fragment to the non-reducing end of another polysaccharide molecule, with the formation of a new glycosidic bond. As there is no net increase in the number of reducing ends in the system, conventional reductometric methods used to assess the activity of glycoside hydrolases are ineffective. Since the PTGs participate in vital processes, such as the elaboration of cell walls in plants and fungi, and are not present in animal cells, they are considered as possible targets for future specific fungicides and herbicides. Biochemical studies of PTGs, as well as the search for their inhibitors, require the availability of convenient and efficient methods for their assay. In this review we briefly describe the principles of methods used to detect and to determine the activity of this important group of enzymes.
Author Contributions
Academic Editor: Bisho Lawaju, Auburn University, Nepal.
Checked for plagiarism: Yes
Review by: Single-blind
Copyright © 2018 Vladimír Farkaš, et al
 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Competing interests
The authors have declared that no competing interests exist.
Citation:
Introduction
Recognition of polysaccharide transglycosylases (PTGs) as important elements of the cell-wall building machinery in plants and fungi has stimulated studies on different aspects of this group of enzymes1,2,3. The PTGs are located in the cell walls, where they participate in cell wall formation and remodeling during growth. Their action enables the inclusion of new cell wall material into the pre-existing wall structure, as well as cell wall extension and modification in the process of growth and maturation. Unlike the glycosyl transferases of Leloir type involved in the biosynthesis of polysaccharides from activated sugars, i.e. from nucleoside-diphosphate sugars, PTGs utilize the energy released by loosening the pre-existing glycoside bond. Since polysaccharide-based cell walls are absent in mammalian cells, it has been proposed that the PTGs might serve as targets for the development of specific fungicides and herbicides4,5,6. Biochemical studies of PTGs, as well as screening for their inhibitors, require the availability of convenient and practical high-throughput assay methods.
In general, the reaction catalyzed by PTGs can be written schematically as follows: (Figure 1)
In the scheme, OOO and ooo are monomer glycosyl units of the donor and of the acceptor, respectively, Ø stands for reducing-end glycosyl unit and (*) is the appropriate label. Products of the transglycosylation reaction are hybrid molecules consisting of a portion of the donor molecule linked to the non-reducing end of the acceptor molecule (P1) and the shortened part of the donor (P2). In cases in which suitable acceptors are not available, some PTGs can transfer the truncated donor chain to a molecule of water, i.e. they can act as hydrolases. When the donor and the acceptor are molecules of the same type, the reaction is called homotransglycosylation; when they are different, we speak about heterotransglycosylation. The molecular size of transglycosylation products depends on the preferred site of attack of the transglycosylase on the donor molecule. Thus, an attack near the non-reducing end (case A, Figure 1) produces transglycosylation products of low relative molecular weight (Mr), whereas splitting the donor molecule close to the reducing end yields products of high Mr (case C) and enzymes attacking the donor molecule stochastically along its length (case B) generate products with a whole range of molecular sizes7.
Figure 1 .Scheme of the transglycosylation reaction. A, B, and C are possible sites of enzyme attack along the polysaccharide main chain.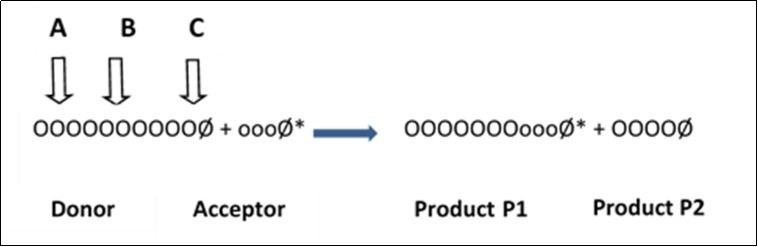
According to Cleland´s nomenclature of enzymatic two-substrate reactions8, the reactions catalyzed by PTGs are of Ping-Pong Bi-Bi type. Accordingly, in the first step the donor molecule is docked to the enzyme active site, the attacked glycosidic bond in the donor molecule is broken, and a portion of the donor molecule carrying the reducing end is released as the product P1. The rest of the donor molecule containing the newly formed reducing end forms a transient glycosyl-enzyme intermediate with the enzyme. After the release of P1, the liberated active sub-sites are occupied with the acceptor and the glycosidic link between the acceptor and the rest of the donor molecule is restored9. The transglycosylation reaction involves double inversion of the glycosidic bond at the C1 of the donor substrate, resulting in retention of anomeric configuration of the glycosidic bond. For a detailed explanation of the reaction mechanism, see e.g. Withers10.
On the basis of their basic principles, PTG assays may be divided into several categories:
A. Assays using labeled acceptors
B. Colorimetric assays
C. Viscometric assays
Assays using Labeled Acceptors
The principle of this type of assay best explains the scheme in Figure 1. Low-Mr oligosaccharides carrying a radioactive or fluorescent label at their reducing ends are used as the acceptors. If the enzyme attacks the donor molecule stochastically along the whole length of the polysaccharide chain, a whole range of molecular sizes of the labeled product is generated7. The accuracy of the assay depends on the effectivity of separation of the labeled products from the labeled acceptors. Their separation is facilitated by the fact that the molecular size of the hybrid products is always higher than that of the acceptors.
There are several ways in which transglycosylation products of different sizes can be separated and quantified:
Sorption.
The radiometric assay designed by Fry et al.2 was historically the first method used to determine the transglycosylation activity of plant xyloglucan endotransglycosylase (XET). In the assay, xyloglucan is the donor and 3H-labeled xyloglucan-derived oligosaccharides (XGOS) or their alditols are used as the acceptors. The key operation in the assay is the separation of the labeled high-Mr transglycosylation products from the unused labeled acceptors. In the case of XET, the ability of xyloglucan to bind firmly to cellulose via hydrogen bonds has been exploited. After the reaction, the assay mixtures are blotted on filter paper and thoroughly washed with water. The radioactivity retained on the paper is taken as the measure of transglycosylase activity. In an alternative method, the so-called “dot-blot” method, xyloglucan and xyloglucan oligosaccharides fluorescently labeled with sulforhodamine (XGOS-SR) were infiltrated into the paper matrix and after drying, the enzyme solution was spotted on the paper and incubated under humid conditions11. After the reaction, the paper was washed to remove unreacted oligosaccharides, leaving fluorescent spots of the xyloglucan‐sulforhodamine reaction product hydrogen‐bonded to the paper. The “dot-blot” method was further improved by arranging the dots on filter paper into the standard 96-well ELISA microplate format and by using the fluorescent microplate reader to quantify the fluorescence12. As an alternative to cellulose matrix for blotting, supports of different kinds, e.g. nitrocellulose13 or glass-fiber paper6, can be used.
The ability of transglycosylases to react with the donor substrates attached non- covalently to solid surfaces was exploited in the design of polysaccharide microarray (or glycochip). Using a robotic DNA spotter, microgram quantities of different polysaccharide donors were printed on the nitrocellulose-covered slides and the microchip was incubated with XET and various fluorescently labeled acceptors14. After washing the slides with 66% ethanol, the fluorescent spots on the microchip revealed the presence of transglycosylase activity. The donors and the acceptors can be freely combined, and for this reason the method can be especially useful when searching for new types of transglycosylating activities.
Here it should be mentioned that the use of fluorescent tags has several advantages over the radiometric ones. Firstly, the investigators do not need to worry about following the special safety regulations that apply to work with radioisotopes. Secondly, fluorescent labels, as well as the instruments required for their detection, are much cheaper than radioactive compounds and the appropriate instrumentation. Third, fluorescent derivatives of saccharides can easily be prepared15. The fluorimetric assays are in principle more sensitive than the radiometric ones. The reason for this is that from the radioactively labeled molecules present in the sample, only a small portion of atoms disintegrates per unit time, whereas in the fluorescently labeled substrate, all the molecules are absorbing and emitting photons concurrently. It should be kept in mind, however, that the sensitivity of the fluorescent methods is influenced by the type of fluorescent label used. The general rule is that the fluorescent tag should be neutral in relation to the enzyme and that the quantum yield, i.e. the ratio between the number of photons emitted and photons absorbed should be close to one. Our previous experience with nasturtium XET showed that sulforhodamine and fluorescein tags are superior to other fluorescent labels tested16.
Chromatography.
Chromatographic methods are universally suitable for separation of individual components from the reaction mixtures. They are especially useful in cases in which molecular characterization of particular product species is required, or when the unreacted labeled acceptors adhere strongly to the absorbent matrix and are difficult to wash out. Size exclusion chromatography is best method to separate large neutral carbohydrate molecules from the small ones. An example of application of HPAEC (High performance anion-exchange chromatography) is the separation and analysis of the transfer products of β-(1,3)-glucanosyltransferase from Aspergillus fumigatus17. Similarly, Nishitani and Tominaga3 as well as Zemková et al.18 measured the activity of xyloglucan endotransglycosylase by resolving the incubation mixtures by size-exclusion HPLC (High performance liquid chromatography, Figure 2). Thin layer chromatography (TLC) on silica gel plates proved to be effective in the resolution and isolation of small-Mr transglycosylation products for determination of their structure by mass spectrometry19.
Figure 2 .Time course of transglycosylation between xyloglucan (donor) and XGOS-SR (acceptors) catalyzed by XET from nasturtium (Tropaeolum majus), as followed by size-exclusion HPLC. Left, fluorescence record of resolution of samples taken at the indicated times of reaction; As the first elute from the column large polysaccharide molecules together with the products of transglycosylation. The second peak corresponds to unused oligosaccharide acceptors XGOS-SR. Right, integrated areas of peaks corresponding to high-Mr transglycosylation products eluting with retention times between 5 and 18 min.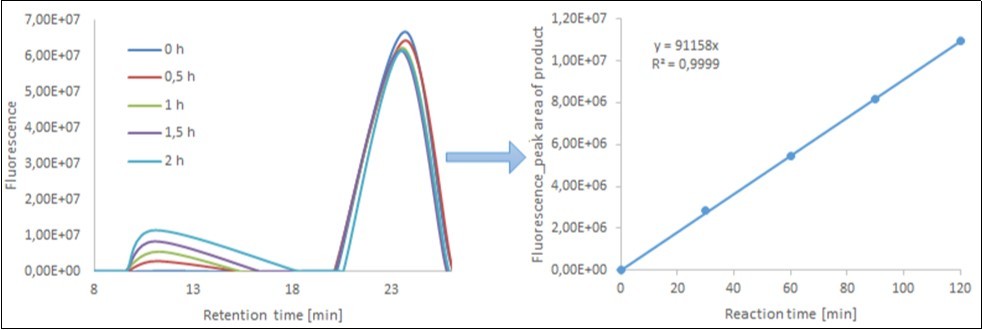
Precipitation.
An alternative means to separate high-Mr products from the oligosaccharide acceptors is the precipitation of the polymer from the reaction mixture with an organic solvent1. The low-Mr products and the unused oligosaccharide acceptors remain in the supernatant, while the radioactively labeled donor:acceptor conjugates are in the precipitate and their radioactivity is determined. The method is quite time consuming because it involves repeated washing and transferring of the precipitated products into sample vials before counting.
Precipitation of the high-Mr transfer products with 66% ethanol is also effective in the detection of active XET species after electrophoresis or isoelectric focusing in polyacrylamide gels by a zymogram technique. After the focusing, the separating gel is overlaid and incubated with a detection gel containing XET substrates: tamarind-seed xyloglucan as the glycosyl donor and sulphorhodamine-labeled xyloglucan-derived oligosaccharides (XGOS-SRs) as the acceptors. By transglycosylation, the xyloglucan in the detection gel becomes labeled. By soaking the detection gel in 66% ethanol, the labeled polymer precipitates and becomes immobilized within the gel matrix, while the unused oligosaccharide acceptors are washed out. The gel is then viewed and photographed under ultraviolet light20. The labeling of polysaccharide donors by the fluorescent acceptors has also been used for the in situ localization of transglycosylase activity in single cells21 as well as in plant organs22, 23, 18. The yeast cells or the plant organs are fed for different periods with fluorescently labeled oligosaccharides, and after their removal by washing with organic solvents, the specimens are observed under a fluorescent microscope. The fluorescence indicates the presence of transglycosylase activity.
Colorimetric Assay
The colorimetric method for measuring transglycosylase activity was specifically designed for XET24. The method exploits the ability of high-Mr xyloglucan to form a blue-green colored complex with iodine. In the presence of native XGOS, the transglycosylase reaction is accelerated, the average Mr of the xyloglucan decreases and the color intensity of the XG:iodine complex is diminished. The transglycosylase activity is then calculated on the basis of the difference of coloration between samples incubated with XGOS added and without additions.
Viscometric Assays
As was already mentioned, the stochastic mode of action of XET on the molecule of xyloglucan7 and the transfer of portions of xyloglucan molecules to small oligosaccharide acceptors cause a decrease in the average molecular weight of the DPS and, consequently, a decrease in viscosity of the solution. The observation that added XGOS significantly stimulated the rate of viscosity decrease of xyloglucan solutions catalyzed by XET, and that the stimulation depended on the concentration of added XGOS, indicated that XGOS behave in the reaction not solely as the product but rather as a second substrate25. The stimulatory effect of added oligosaccharides on the rate of polymer degradation may be used to distinguish endotransglycosylases from endohydrolases, since the “classical” endoglycanases are not stimulated by added oligosaccharides. The viscometric assay was also used to prove the heterotransglycosylation activities between xyloglucan and xylan, and xyloglucan and galactomannan in vitro in extracts from charophyte green algae26.
High Throughputs.
Specific properties of fungal polysaccharide transglycosylases, such as their participation in fungal and plant cell wall formation and modification, make them attractive targets for future specific antifungals and/or herbicides. One of the possible approaches to the search for specific inhibitors of transglycosylases is the screening of a great number of candidate compounds using high-throughput (HTP) techniques.
The “high-throughput” techniques use the common multi-well ELISA (Enzyme-Linked Immunosorbent Assay) plates, in which the incubations and all the subsequent steps, such as washing and quantitative determinations, are performed in the same well. Avoiding sample transfers reduces the incidence of errors. Importantly, the HTP methods enable the analysis of multiple samples in parallel, which makes statistic evaluation of the results more precise. To facilitate the GTP assays, we have devised two variants of their high-throughput assay. Both variants are based on the fluorescent assay described above, and in both methods the key point is retaining the high-Mr fluorescent products of transglycosylation and eliminating the low-Mr oligosaccharide acceptors27.
In the variant called “in-gel assay”, the reaction takes place in a gel sticking to the bottom of the microplate well. The reaction mixture consists of both the polysaccharide donor and the fluorescently labeled oligosaccharide acceptor incorporated into a buffered agar gel. The assay gel is deposited at the bottom of a microplate well and the enzyme is added to start the reaction (Figure 3). The reaction is terminated by addition of 66% acidified ethanol. Ethanol precipitates the polysaccharide molecules, both labeled and unlabeled, while the unused labeled oligosaccharides are removed by washing. The fluorescence retained in the agar corresponds to the transglycosylase activity. The advantage of the in-gel assay is its simplicity and high sensitivity due to the three-dimensional reaction space; the disadvantage is the need for prolonged and repeated washings in order to obtain a low fluorescence background.
Figure 3.Scheme of the transglycosylase in-gel assay. The reaction mixture contains 0.05% donor polysaccharide, 4 µM acceptor oligosaccharide in 1.5 % agar dissolved in 0.05 M citrate-phosphate buffer, pH 3.5 and the enzyme. The reaction is stopped by acidified 66% ethanol and the gel is washed thoroughly with 66% ethanol.
The second HTP variant exploits the capability of the transglycosylase reaction also to proceed with donor polysaccharide adsorbed on the surface of the microplate well. In order to increase the affinity of the sorption of the polysaccharide donors on the microplate polymer, the bottom surface of the wells is pre-coated with nitrocellulose (NC). DPS is then adsorbed on top of the nitrocellulose layer and after addition of acceptor and the enzyme, the reaction proceeds at the interface between the attached DPS and the liquid (Figure 4). At the end of the incubation, the reaction is stopped by addition of 66% ethanol containing 5% HCOOH, the wells are rinsed several times with the same solution and the fluorescence adhering to the wells is determined. A great advantage of both HTP variants of the assay is their versatility concerning the type of transglycosylation reaction assayed, as well as the capacity to handle many samples simultaneously.
Figure 4 .Scheme illustrating steps in the high throughput PTG assay performed in microplate wells coated with nitrocellulose (NC) with adsorbed donor polysaccharide (DPS). For explanation see the text.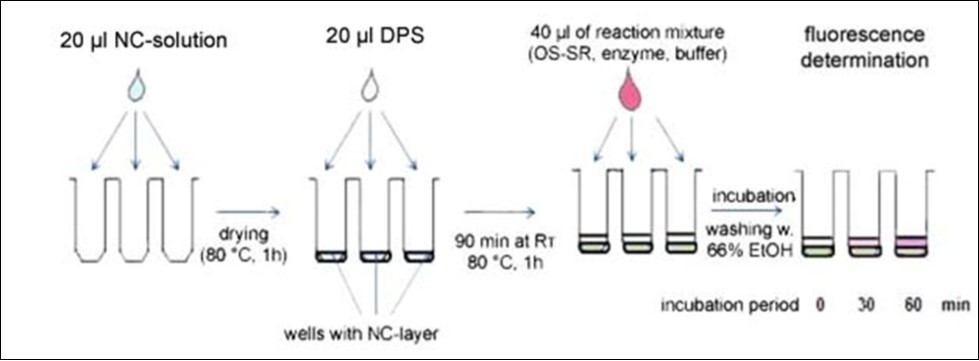
Conclusion
With this review we tried to describe the basic principles on which the currently used PTG assays are built. The key problem is ultimately to accommodate the method to the user´s particular needs and the specific conditions of the experiment. Sometimes at the application of a new method, pitfalls or problems may occur that went unnoticed by the original author. The reasons may be different, often it is the different quality of substrates that are mostly non-commercial and have to be prepared. The combination of assay methods based on different principles may be useful; it can often help to better characterize the properties of the studied enzyme(s).
Acknowledgements
The work for this review was supported in part by the grant no. 2/0058/16 from the Grant Agency for Science VEGA (Slovakia) to E. S. We thank Michael J. Bailey for correcting the English style.
References
- 1.Farkaš V, Sulová Z, Stratilová E, Hanna R, Maclachlan G. (1992) Cleavage of xyloglucan by nasturtium seed xyloglucanase and transglycosylation to xyloglucan subunit oligosaccharides. , Arch. Biochem. Biophys 298(2), 365-370.
- 2.S C Fry, R C Smith, K F Renwick, D J Martin, S K Hodge et al. (1992) Xyloglucan endotransglycosylase, a new wall-loosening enzyme activity from plants.Biochem.J;282(Pt3). 821.
- 3.Nishitani K, Tominaga R. (1992) Endo-xyloglucan transferase, a novel class of glycosyltransferase that catalyzes transfer of a segment of xyloglucan molecule to another xyloglucan molecule. , J. Biol. Chem 267, 21058-21064.
- 4.J P Latgé. (2007) The cell wall: a carbohydrate armour for the fungal cell. , Mol. Microbiol 66(2), 279-290.
- 5.Delso I, Valero-Gonzales J, Gomollon-Bel F, Castro-Lopez J, Fang W. (2018) Inhibitors against cell wall remodelling enzymes. , ChemMedChem 13, 128-132.
- 6.Chormova D, Franková L, Defries A, S R Cutler, S C Fry. (2015) Discovery of small molecule inhibitors of xyloglucan endotransglucosylase (XET) activity by high-throughput screening. , Phytochemistry 117, 220-236.
- 7.N M Steele, Sulová Z, Campbell P, Braam J, Farkas V et al. (2001) Ten isoenzymes of xyloglucan endotransglycosylase from plant cell walls select and cleave the donor substrate stochastically.Biochem.J,355(Pt3). 671.
- 8.Cleland W. (1963) The kinetics of enzyme-catalyzed reactions with two or more substrates or products: I. Nomenclature and rate equations. , Biochim. Biophys. CES. Acta (BBA)-Specialized Section on Enzymological Subjects 67, 104-137.
- 9.Baran R, Sulová Z, Stratilová E, Farkaš V. (2000) Ping-pong character of nasturtium-seed xyloglucan endotransglycosylase (XET) reaction. , Gen. Physiol. Biophys 19(4), 427-440.
- 10.S C Fry. (1997) Novel ‘dot-blot’ assays for glycosyltransferases and glycosylhydrolases: optimisation for xyloglucan endotransglycosylase (XET) activity. , Plant J 11, 1141-1150.
- 11.Mazáň M, Ragni E, Popolo L, Farkaš V. (2011) Catalytic properties of the Gas family β-(1, 3)-glucanosyltransferases active in fungal cell-wall biogenesis as determined by a novel fluorescent assay. , Biochem. J 438(2), 275-282.
- 12.Stratilová E, Ait-Mohand F, Řehulka P, Garajová S, Flodrová D et al. (2010) Xyloglucan endotransglycosylases (XETs) from germinating nasturtium (Tropaeolum majus) seeds: isolation and characterization of the major form. , Plant Physiol. Biochem 48(4), 207-215.
- 13.Kosík O, R P Auburn, Russell S, Stratilová E, Garajová S et al. (2010) Polysaccharide microarrays for high-throughput screening of transglycosylase activities in plant extracts. , Glycoconj. J 27, 79-87.
- 14.Kosík O, Farkaš V. (2008) One-pot fluorescent labeling of xyloglucan oligosaccharides with sulforhodamine. , Anal. Biochem 375(2), 232-236.
- 15.Kosík O, Garajová S, Matulová M, Řehulka P, Stratilová E et al. (2011) Effect of the label of oligosaccharide acceptors on the kinetic parameters of nasturtium seed xyloglucan endotransglycosylase (XET). , Carbohydr. Res 346(2), 357-361.
- 16.R P Hartland, Fontaine T, J P Debeaupuis, Simenel C, Delepierre M et al. (1996) A novel β-(1,3)-glucanosyltransferase from the cell wall of Aspergillus fumigatus. , J. Biol. Chem 271(43), 26843-26849.
- 17.Zemková Z, Garajová S, Flodrová D, Řehulka P, Zelko I et al. (2012) Incorporation of β-(1,6)-linked glucooligosaccharides (pustulooligosaccharides) into plant cell wall structures. , Chem. Papers 66, 814-820.
- 18.Kováčová K, Degani G, Stratilová E, Farkaš V, Popolo L. (2015) Catalytic properties of Phr family members of cell wall glucan remodeling enzymes: implications for the adaptation of Candida albicans to ambient pH. doi: 10.1093/femsy/fou011. , FEMS Yeast Res 15(2).
- 19.Farkaš V, Ait-Mohand F, Stratilová E. (2005) Sensitive detection of transglycosylating activity of xyloglucan endotransglycosylase/hydrolase (XTH) after isoelectric focusing in polyacrylamide gels. , Plant Physiol. Biochem 43(5), 431-435.
- 20.Cabib E, Farkas V, Kosík O, Blanco N, Arroyo J et al. (2008) Assembly of the yeast cell wall Crh1p and Crh2p act as transglycosylases in vivo and in vitro. , J. Biol Chem 283(44), 29859-29872.
- 21.Vissenberg K, I M Martinez-Vilchez, J P Verbelen, J G Miller, S C Fry. (2000) In vivo colocalization of xyloglucan endotransglycosylase activity and its donor substrate in the elongation zone of Arabidopsis roots. , The Plant Cell 12, 1229-1237.
- 22.S G Withers. (2001) Mechanisms of glycosyl transferases and hydrolases. , Carbohydr. Polymers 44(4), 325-337.
- 23.F M Ibatullin, Banasiak A, M J Baumann, Greffe L, Takahashi J et al. (2009) A real-time fluorogenic assay for the visualization of glycoside hydrolase activity in planta. , Plant Physiol 151, 1741-1750.
- 24.Sulová Z, Lednická M, Farkaš V. (1995) A colorimetric assay for xyloglucan-endotransglycosylase from germinating seeds. , Anal. Biochem 229(1), 80-85.
- 25.Farkas V, Maclachlan G. (1988) Stimulation of pea 1, 4-β-glucanase activity by oligosaccharides derived from xyloglucan. , Carbohydr. Res 184, 213-219.
Cited by (2)
This article has been cited by 2 scholarly works according to:
Citing Articles:
Food Science and Biotechnology (2023) Crossref
Food Science and Biotechnology (2023) OpenAlex
I. Siziya, Jong-Hyun Jung, Myung-Ji Seo, Min Lim, D. Seo - Food Science and Biotechnology (2023) Semantic Scholar
