Abstract
Objective:
MicroRNAs are involved in the onset, progression and dissemination of esophageal cancer, and they may be useful as prognostic biomarkers. This study aims to evaluate the relation of miR-21 expression and the prognosis of esophageal cancer patients.
Methods:
In this study, a meta-analysis is performed by searching PubMed, Science Direct databases, and Cochrane Library. Data are extracted from studies evaluating survival of esophageal cancer patients with either high or low miR-21 expression. Pooled hazard ratios (HRs) and 95% confidence intervals (CI) are calculated.
Results:
A total of 579 cases of esophageal cancer from five studies are involved for this global meta-analysis. The HR of survival of patients with high miR-21 expression is 1.47 (95% CI: 1.12–1.91; p<0.01) as compared with those with low expression.
Conclusions: miR-21 may be a predictor for survival of esophageal cancer patients.
Author Contributions
Academic Editor: Ying-Chu Lin, Kaohsiung Medical University, Taiwan
Checked for plagiarism: Yes
Review by: Single-blind
Copyright © 2016 Wei Zhou, et al.
 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Competing interests
The authors have declared that no competing interests exist.
Citation:
Introduction
Esophageal cancer (EC) is the eighth most common cancer in the world, affecting approximately 500,000 individuals worldwide each year 1, 2. EC consists of two predominant histological subtypes, squamous cell carcinoma and adenocarcinoma 3. Esophageal squamous cell carcinoma accounts for more than 90% of esophageal carcinomas in Asian countries, while esophageal adenocarcinoma affects white populations predominantly. The biology of EC was of aggressive local invasion, early metastasis, and resistance to chemotherapy. Despite improvements in detection, surgical resection, and adjuvant therapy, the prognosis for long-term survival of EC remains poor 4.
MicroRNAs (miRNAs) are small non-protein coding RNAs that function as negative gene regulators at the post-transcriptional level 5, 6. The mature miRNA destabilizes the target mRNA transcript or blocks its translation or both, thus affecting fundamental biological processes, such as cell proliferation, apoptosis, and differentiation. A growing amount of evidences proves that miRNAs are involved in the onset, progression and dissemination of EC, and they may be useful as both diagnostic biomarkers and therapeutic targets 7, 8, 9. miR-21, an oncomiR, is significantly up-regulated in almost all types of cancers, including breast cancer, EC, head and neck squamous cell carcinoma (HNSCC), etc 10, 11. High expression of miR-21 is correlated with poor overall survival in HNSCC, suggesting that it may serve as a prognostic marker for human cancers 12, 13.
In this study, we have firstly performed a meta-analysis to evaluate the relationship between miR-21 expression level and the survival of EC patients. We have also discussed the current status of using miR-21 as a prognostic marker and therapeutic target in EC.
Material and Methods
Study selection
This meta-analysis is carried our following the guidelines of the meta-analysis of Observational Studies in Epidemiology group (MOOSE) 12, 14. A search is applied to the following electronic databases: EMBASE, PubMed, Cochrane Library and Science Direct database from 1980 to Oct 2013. The following key search terms are used: ‘miR-21 and cancer’ and ‘microRNA, cancer and prognosis’. The search is carried out on studies with human subjects, without restriction on language. The reference lists were also reviewed manually to identify additional studies. When overlapping data of the same patient population are included in more than one publication, only the most recent or complete study is used in this study.
Inclusion/exclusion criteria
The following criteria were used to include published studies: (1) the study has to investigate the patients with EC; (2) the study has to detect the expression of miR-21 in tissue; (3) the study has to detect the association between the expression levels of miR-21 and survival; (4) the follow-up time is more than 2 years; (5) the study is published as a full paper from January 1980 to Oct 2013.
The following criteria were used to exclude published studies: (1) letters, (2) lack of key information, including HR and 95% CI of each study.
Quality assessment
The quality of all the studies is systematically evaluated according to a critical review checklist of the Dutch Cochrane Centre proposed by MOOSE 15. The following key points are clearly checked: patient population, origin of country, tumor type, method of detecting miR-21, outcome, survival analysis and sufficient period of follow-up. Any study without mentioning these points is excluded so as not to compromise the quality of this meta-analysis.
All reviewers are physician investigators with systematic review experience. The information such as titles, abstracts, full texts and reference lists of all of the identified reports are carefully identified in duplicate by two investigators (Liu and Zhang). These extracted articles are double checked by Zhou and Hong. All the disagreement is resolved by consensus. The references from the relevant literature, including all the identified original articles, reviews, comments and editorials, are reviewed manually. The additional information and the data needed for the meta-analytic calculations are obtained by emailing to the authors.
Data extraction
The following data are extracted from each study in a predefined form (Table 1): first author's name, country of origin, total number of cases (N), method of detecting miR-21, tumor type, follow-up time, etc. An HR of > 1 represents an increase likelihood of death. If only survival curves are available, data are extracted from the graphical survival plots according to the previously described method 14, 15.
Statistical analysis
Statistics are conducted as described previously 15. HR and 95% CI of each study are calculated. Pooled HR is calculated using a random-effects model. A P value less than 0.05 is considered statistically significant. All analyses were conducted using ‘STATA: Data Analysis and Statistical Software’ V10.1 (http:// www.stata.com; Stata Corporation, College Station, TX). A random effect model was applied if heterogeneity existed (P<0.05), while the fixed effect model was utilized in the absence of between-study heterogeneity (P≥0.05). Potential publication bias was assessed by Begg’s funnel plots and Egger’s regression test. Our work focused on identifying the consistence of differently expressed miRNAs, which is of great value in improving the diagnostics, therapeutics and prognosis in carcinoma.
Results
Search results:
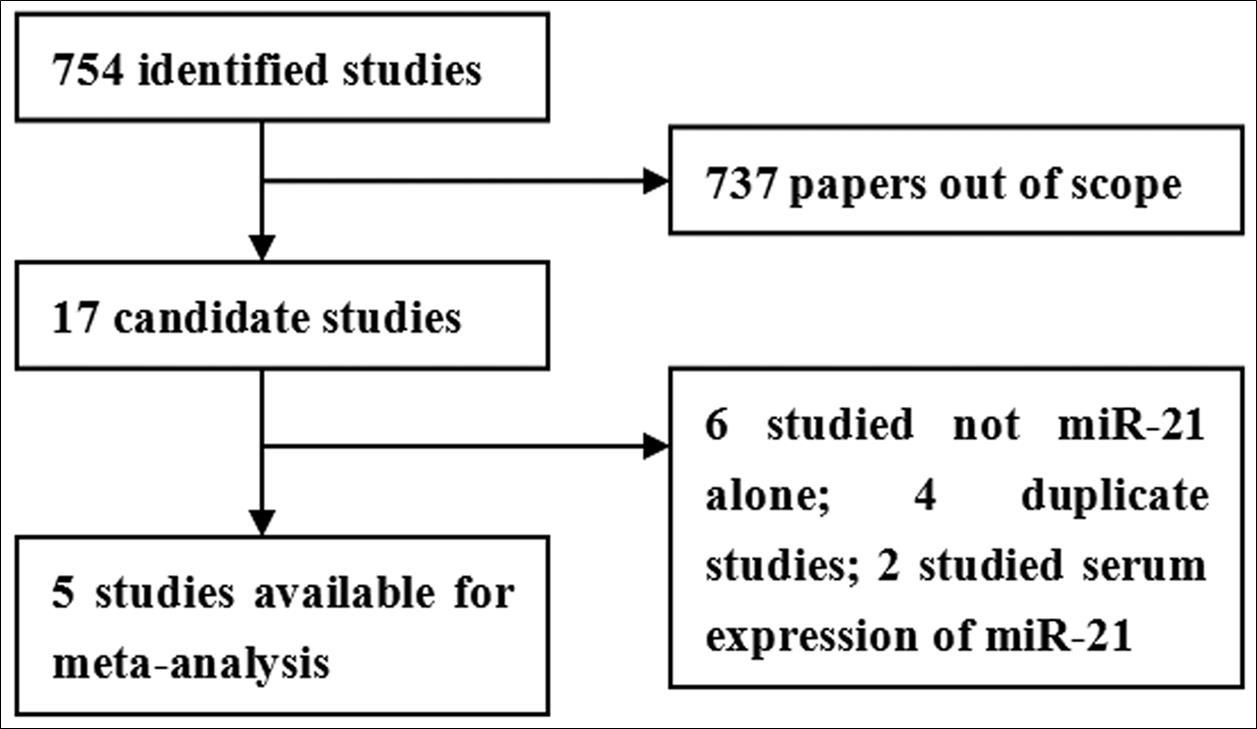
As shown in Figure 1, seven hundred and fifty four records are identified. After review of title and abstracts, seven hundred and thirty seven studies are excluded due to their irrelevance to the current analysis. Furthermore, twelve candidate studies are excluded because of the following reasons: 6 studies without detecting miR-21 alone; 4 duplicate studies; 2 studies focusing on serum expression of miR-21 16, 17. Therefore, the final meta-analysis was performed on the basis of the remaining 5 studies 18, 19, 20, 21, 22. The main features of eligible studies are summarized in Table 1.
High expression of miR-21 predicts poor survival:
The data of 5 studies from the United States, China and Japan were collected, including a total of 579 cases of EC patients. The methods of detecting miR-21 expression are in situ hybridization, quantitative real-time PCR (qRT-PCR) and stem–loop RT-PCR. The pooled HR is 1.47, and 95% CI is 1.12–1.91. As shown in Figure 2, high expression level of miR-21 significantly predicts poor survival (P < 0.01). There was no evidence for publication bias on the funnel plot (data not shown).
Figure 2.Forest plot for the association between high miR-21 expression and survival of patients with esophageal cancer.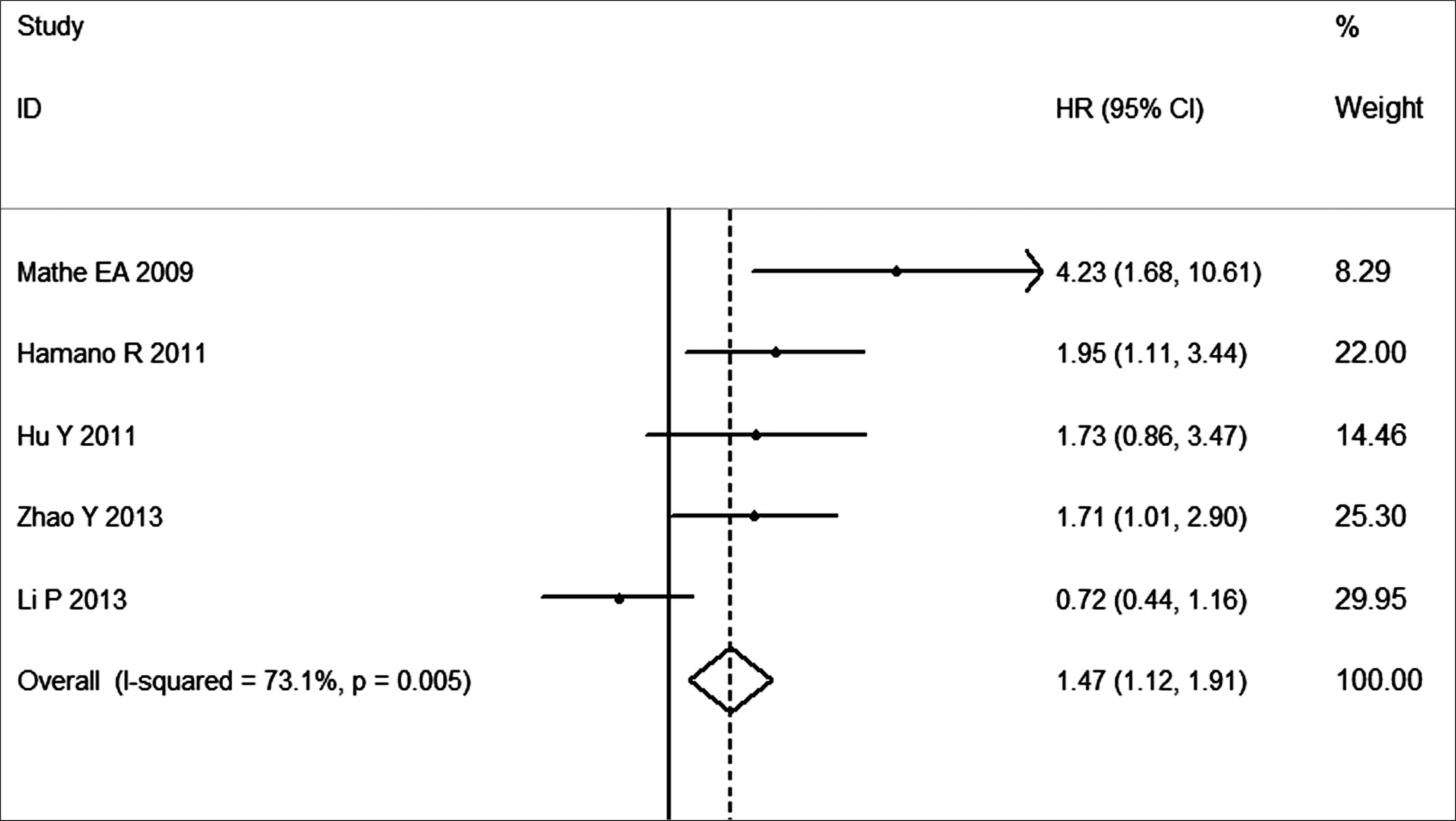
Discussion
miRNAs are critical regulators of transcriptional and post-transcriptional gene silencing, which were involved in multiple developmental processes in many organisms 23, 24. miRNAs are aberrantly expressed or mutated in human cancer, indicating that they may function as a novel class of oncogenes or tumor suppressor genes. miR-21is overexpressed in most tumor types, and acts as an oncogene. In a variety of cancer cell lines, up-regulation of miR-21 leads to increased cell proliferation, invasion, and migration by regulation of cell cycle and apoptosis 10, 11. Due to its ubiquitous role in various biological processes, the interest in miR-21 has dramatically increased during recent years, especially in cancer.
In this study, totally 5 studies with the follow-up time of more than 2 years were involved. Hamano et al 19 have found that overexpression of miR-200c could induce chemoresistance in EC through activation of the Akt signaling pathway . Kaplan–Meier curves were used to detect the overall survival rates of 98 patients with EC who received preoperative chemotherapy followed by surgery. High expression levels of miR-200c and miR-21 and low expression levels of miR-145 correlated significantly with shorter overall survival. Mathe et al 18 have detected the expression of miR-21 in 170 cases of EC cancerous and adjacent noncancerous tissues. Elevated expression of miR-21 in noncancerous tissue was associated with worse prognosis. Li et al 21 have found that miR-21 was significantly overexpressed in EC tissues and cell lines. There was significant difference in the median survival (13 vs. 19 months) between patients with miR-21 overexpression (>5-folds) and those patients with miR-21 expression (<5-folds). Zhao et al 22 have found that the expression patterns of miR-21, miR-181b and miR-146b could be used as prognostic classifiers for patients with EC. High expression of miR-21 in EC tissue was significantly associated with worse survival.
This systemic review and meta-analysis shows that elevated miR-21 expression may predict poor survival in EC patients. In addition, miR-21 may be used as a therapeutic target in EC. miR-21 has been identified as one of the first chemoresistance related miRNAs 13. As for patients with EC who have received preoperative chemotherapy followed by surgery, overexpression of miR-21 correlates significantly with shortened overall duration of survival. Inactivating endogenous miR-21 is considered as a new strategy that may increase the sensitivity of EC cells to anticancer drugs. So far, the exact molecular mechanism of miR-21 deregulation and its roles in development of EC are unclear.
his meta-analysis has several limitations. First, the pooled HR is calculated on the basis of 5 studies with a small sample size of 579 patients. Second, the cut-off definition of miR-21 is different in each study. Third, the expression of miR-21 is detected in tissue samples, instead of serum.
Table 1. Summary of studies reporting miR-21 expression and survival of esophageal cancer| First author (year) | Country | N | miR-21 assay | Tumor type | Follow-up, months | Survival analysis |
| Mathe EA (2009) 18 | Japan, USA | 69 | qRT-PCR | ESCC | 60 | Kaplan-Meier analyses |
| Hamano R (2011) 19 | Japan | 98 | qRT-PCR | ESCC | 96.7 | Kaplan-Meier analyses |
| Hu Y (2011) 20 | USA | 158 | In situ hybridization | EA | 256.43 | Kaplan-Meier analyses |
| Li P (2013) 21 | China | 76 | Stem–loop RT-PCR | ESCC | 30 | Kaplan-Meier analyses |
| Zhao Y (2013) 22 | China | 178 | qRT-PCR | ESCC | 40 | Kaplan-Meier analyses |
Several limitations in this meta-analysis should be acknowledged. First, in reference 18, there was a correlation between high expression of miR-21 gene in non-cancerous tissues and poor survival of patients with esophageal cancer. Due to the lack of the related literature, further studies need to be conducted in the future. Second, no significant association was found between miR-21 gene expression and prognosis of esophageal cancer using the data in cancerous tissue. Hence, a precise analysis should be performed. Third, the other 4 studies were just focusing on the high expression of miR-21 gene in cancerous tissues, which did not involve survival rate. Forth, it should be noted that the carcinogenesis for high expression of miR-21 protein in cancerous tissue was highly possible different that (high expression of miR-21 protein) in non-cancerous tissue. Therefore, a single high expression of microRNA in cancerous tissue might not be enough to explain cancer prognosis. Finally, the patients with esophageal adenocarcinoma were recruited into the study of reference 20 on table 1, which is one of 5 studies included into present study. It needs to be claimed that the mechanisms of cancer development might be different between esophageal squamous cell carcinoma and esophageal adenocarcinoma.
In conclusion, miR-21 expression might be an independent predictor for outcome of EC patients. High expression of miR-21 is significantly associated with poor survival of EC patients. Further analysis of miR-21 targets and downstream functions are necessary to fully understand the role of miR-21 in EC.
Acknowledgements
This study was supported in part by grants from the National Natural Scientific Foundation of China (81100714, 81171923). There is no conflict of interest.
References
- 2.Hong L, Han Y.Lu Q,et al.Drug resistance-related microRNAs in esophageal cancer. (2012)Expert Opin Biol Ther.12,1487-1494.
- 4.DR Nieman, JH Peters.Treatment strategies for esophageal cancer. (2013)Gastroenterol Clin North Am.42,187-197.
- 5.Gu J, Wang Y, Wu X.MicroRNA in the pathogenesis and prognosis of esophageal cancer. (2013)Curr Pharm Des.19,1292-1300.
- 6.Fassan M, Baffa R, Kiss A.MicroRNA dysregulation in esophageal neoplasia: the biological rationale for novel therapeutic options. (2013)Curr Pharm Des.19,1236-1241.
- 7.GC Mayne, DJ Hussey, DI Watson.MicroRNAs and esophageal cancer--implications for pathogenesis and therapy. (2013)Curr Pharm Des.19,1211-1226.
- 10.Kumarswamy R, Volkmann I, Thum T.Regulation and function of miRNA-21 in health and disease. 2011-8.
- 12.Fu X, Han Y.Wu Y,et al.Prognostic role of microRNA-21 in various carcinomas: a systematic review and meta-analysis. , (2011)Eur J Clin Invest.41,1245-1253
- 13.Hong L, Han Y.Zhang Y,et al.MicroRNA-21: a therapeutic target for reversing drug resistance in cancer. (2013)Expert Opin Ther Targets.17,1073-1080.
- 14.Hong L, Yang J, Han Y.High expression of miR-210 predicts poor survival in patients with breast cancer: a meta-analysis. 2012-507.
- 15.Hong L, Zhang H, Zhao Q.Relation of excess body weight and survival in patients with esophageal adenocarcinoma: a meta-analysis. (2013)Dis Esophagus.26,623-627.
- 16.Tanaka K, Miyata H, Yamasaki.M,et al.Circulating miR-200c Levels Significantly Predict Response to Chemotherapy and Prognosis of Patients Undergoing Neoadjuvant Chemotherapy for Esophageal Cancer. (2013)Ann Surg Oncol.20,607-615.
- 17.Komatsu S, Ichikawa D.Takeshita H,et al.Prognostic impact of circulating miR-21 and miR-375 in plasma of patients with esophageal squamous cell carcinoma. 2012-12.
- 18.EA Mathé, GH Nguyen.Bowman ED,et al.MicroRNA expression in squamous cell carcinoma and adenocarcinoma of the esophagus: associations with survival. (2009)ClinCancer Res. 15-6192.
- 19.Hamano R, Miyata H, Yamasaki.M,et al.Overexpression of miR-200c induces chemoresistance in esophageal cancers mediated through activation of the Akt signaling pathway. (2011)ClinCancer Res. 17-3029.
- 20.Hu Y, AM Correa.Hoque A,et al.Prognostic significance of differentially expressed miRNAs in esophageal cancer. (2011)IntJ Cancer.128,132-143
- 21.Li P, WM Mao, ZG Zheng.. Down-Regulation of PTEN Expression Modulated by Dysregulated miR-21 Contributes to the Progression of Esophageal Cancer. (2013)Digest Dis Sci 58-3483.
- 22.Zhao Y, AJ Schetter.Yang GB,et al.microRNA and inflammatory gene expression as prognostic marker for overall survival in esophageal squamous cell carcinoma. (2013)IntJ Cancer.132,2901-2909
Cited by (1)
- 1.Zhou Wei, Yang Wanli, Yang Jing, Zhu Haijun, Duan Lili, et al, 2021, miR‑483 promotes the development of colorectal cancer by inhibiting the expression level of EI24, Molecular Medicine Reports, 24(2), 10.3892/mmr.2021.12206