Testing Maternal-Fetal Genotype Incompatibility with Mother-Offspring Pair Data
Abstract
Maternal-fetal genotype (MFG) incompatibility has been shown to be strongly associated with a number of genetic diseases. Most current methods for the MFG test rely on parents-offspring trio genotype data and do not apply to mother-offspring pair data where paternal genotype is unavailable. In this paper, we developed a two-stage method for testing the allelic effect of given SNP through subgroup analysis of compatible MFG pairs and further tested the MFG incompatibility effect according to different scenarios determined by the allelic effect. Simulation studies demonstrated that this novel two-stage model is powerful in detecting the MFG incompatibility effect. This method can be implemented through publicly available statistical software, such as SAS, R, etc. We demonstrate with a case-control study of small for gestational age neonates that the method identified a SNP in the IGF2R gene with a significant allelic effect (p = 0.037), and a SNP in the IGF1 gene with a significant MFG incompatibility effect (OR = 0.75 and p = 0.033).
Author Contributions
Academic Editor: Shanzhi Wang, Biochemistry department, Albert Einstein College of Medicine, United States.
Checked for plagiarism: Yes
Review by: Single-blind
Copyright © 2013 Wenjiang J. Fu, et al.
 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Competing interests
The authors have declared that no competing interests exist.
Citation:
Background
Single nucleotide polymorphisms (SNPs) have been shown with growing evidence to be associated with many diseases through polymorphisms at single locus or through haplotype at multiple loci (Zhang et al. 2004, Spinka et al. 2004, Cui et al. 2007).43,37,9. Pediatric studies have demonstrated strong association of phenotypes or childhood diseases with parental genotypes, offspring genotypes or both (Goddard et al. 2007 and references therein)14 . In addition, research using both maternal and perinatal genotype data focusing on the interaction between the maternal and offspring genes has provided further evidence in gene-gene interaction, in particular, the effect of genotype incompatibility between the maternal and fetal genes has been found to play a critical role (Sinsheimer, Elston and Fu 2011). Hence searching for genes of incompatible maternal-fetal genotypes (MFG) is of particular interest.
Studies of the incompatibility of MFG can be traced back to as early as 1960s on hemolytic disease of newborn caused by anti-K antibody (Kulich V, Kout M. 1967)19 and neonatal thrombopenia feto maternal incompatibility in human leukocytes antigen (HL-A) system (Salet et al. 1973a and 1973b)33,34. However, not until 1990s had the MFG received much attention (Hollister et al 1996). Although early studies introduced the MFG concept in human diseases research (Kulich V, Kout M. 1967, Salet et al. 1973a and 1973b, Stubbs, Ritvo and Mason-Brothers 1985)33,34,19,38 , statistical modeling of the MFG was not employed until 2002.
In a pioneering work, a MFG loglinear model was proposed to examine the MFG incompatibility effect by analyzing parents-offspring trio genotype data (Sinsheimer et al 2003) 36and was applied to examine the association with the susceptibility of schizophrenia (Palmer et al. 2002)29. The MFG loglinear model is based on Poisson regression, which divides the samples into different categories of all combinations of parents-offspring trio genotypes. The number of cases in each category is then modeled with loglinear model following a Poisson distribution. Without Hardy-Weinberg equilibrium (HWE) assumption, the expectation of the number of cases in each category was parameterized with corresponding mating rate, mother or child genetic effects as well as the MFG incompatibility effect. The parameters are then estimated through the maximum likelihood approach and the inference is made accordingly.
The initial loglinear model for MFG test was improved in a series of articles. The correlation between siblings and relatives was incorporated (Kraft et al 2004)17. An exact MFG test was proposed to increase the statistical power of the likelihood ratio test by permuting the trio counts to generate a background distribution (Minassian et al 2005)25. The MFG test has so far been applied to study a number of diseases, including RhD and HLA-B for schizophrenia, ABO for autism etc. (Palmer et al 2006, Zandi et al, 2006)30,42. Recently, the MFG test was further extended for nuclear family data (Minassian et al 2006, Childs et al 2008).26,8
The loglinear model for MFG test has been proven to be successful in a number of studies. In general, its requirement of parents-offspring trio genotype data, however, may limit its application to many studies where no paternal genotype data are available, although the statistical expectation-maximization (EM) algorithm or direct maximization of the observed loglikelihood (Misassian 2006, Hsieh 2006)26,30 can be implemented to deal with missing data when a portion of samples have missing paternal or maternal genotypes. Shi et al (2008)35 showed that Mendelian inheritance, random mating and parental allelic exchangeability may be incorporated in the loglinear model simultaneously. Chen, Zheng, and Wilson (2009)7 took a retrospective likelihood approach to achieve the same effect of modeling family information. Recently, Chen et al (2012)6 further studied a semiparametric model to incorporate environmental factors. These models have been demonstrated with consistent and efficient estimation. However, due to the model complexity, the interpretation is not readily understood by many principal investigators and the model is not easy to implement by non-statistical professionals. Hence it is desirable and attractive to have a simple and yet efficient model to provide an easy-to-understand-and- implement procedure using currently available statistical software.
The fact that many perinatal studies with mother-offspring pairs do not have paternal genotype makes it desirable to extend the MFG test from parents-offspring trio genotype data to mother-offspring pair genotype data. For this purpose, we extended the MFG test and developed a two-stage logistic regression model. This model conducts analysis in two steps. In the first step, it tests allelic effect of given SNP through a subgroup analysis using only mother-offspring pairs that had compatible MFG. It requires a moderately large sample size which allows subgroup analysis with sufficient power for testing the allelic effect and leads to several scenarios according to the allelic effect. In the second step, the MFG incompatibility effect is tested using all mother-offspring pairs through a logistic regression model accommodating the allelic effect with different scenarios. This two-stage model is different from the ones proposed by Li et al. (2009)21 and Ainsworth et al. (2011)1. It enjoys meaningful interpretation of logistic regression and the allelic effects of SNPs, and can be implemented with common statistical software, such as SAS, R, etc. Statistical simulation studies demonstrate that this two-stage model achieves sufficient power to detect the MFG incompatibility effect. The method is also illustrated by an application to a case-control study of small for gestational age (SGA) neonates.
Materials and Method
Data
SGA Dataset
A total of 991 mother-offspring pairs (406 SGA cases and 585 controls) were included in this case-control study conducted in Chile. SGA was defined as birth weight below 10th percentile for the corresponding gestational age Kramer 198718. Clinical and socio-demographic data were summarized in Table S1 (Supplementary Material), which shows no statistical significance between the two groups in maternal age, maternal BMI, smoking status and baby’s gender. Parity was marginally significant with p=0.048. The control group had a larger mean gestational age and larger mean birth weight as expected.
SNP Data
Template DNA was obtained for genotyping through whole genome amplification of genomic DNA extracted from blood samples of patients following an automated DNA isolation protocol (BioRobot 9604, Qiagen, Valencia, CA, USA). Genotyping of single nucleotide polymorphisms (SNPs) was carried out using the massARRAYTM System (Sequenom Inc. San Diego, CA, USA) by the high-throughput genotyping facility at Genaissance Inc (New Haven, CT, USA). SNP genotyping quality control was conducted through genotyping consistency of repeated samples. More than 1300 SNPs were genotyped in this candidate gene based case-control study for several phenotypes of pregnancy complication, including preeclampsia, SGA, etc. We focus on the IGF-I and IGF-II genes and their receptors only in this study.
A total of 44 SNPs were genotyped in IGF-I, IGF-II, IGF1R and IGF2R genes. Among them, 5 SNPs were found to be homozygous in the study population and 13 SNPs had one minor allele of frequency less than 0.01 and thus were excluded. HWE was examined in the combined mother and offspring control population and all of the 26 remaining SNPs satisfied the HWE with a cut-off p-value > 0.0001. Ten SNPs out of these 26 remaining SNPs were found to have one minor allele with frequency equal to or smaller than 0.1 (homozygous genotype probability ≤ 0.01) and were excluded from the MFG incompatibility test. Thus the MFG incompatibility test was conducted on the remaining 16 SNPs.
Methods
We develop a two-stage model for testing the MFG incompatibility, and further conduct a simulation study.
The Two-Stage Method for MFG Incompatibility
Step 1. Determination of allelic effect
Step 1 determines the allelic effect between the two alleles and its scenario at each SNP. We first examine the minor allele frequency and the HWE for each SNP. Only those SNPs having a minor allele frequency (p ≥ 0.01) and satisfying the HWE are kept for further study on the examination of allelic effect. The HWE is examined with a chi-square test in the combined mother and offspring control populations with a cut-off value of chi-square test p-value > 0.0001.
To study the gene-gene interaction between the mother and her offspring at a given SNP, a composite SNP genotype of the mother and offspring pair is derived by combining the two. For example, if the mother genotype at a given SNP is ‘AA’ and the offspring genotype is ‘AB’, the composite genotype is ‘AA_AB’. There are seven possible composite genotypes, see Table 1 for details. Among them, three are compatible: ‘AA_AA’, ‘AB_AB’ and ‘BB_BB’, and the rest are incompatible.
Table 1. Composite genotypes of the mother-offspring pair| Mother genotype | Offspring genotype | ||
| AA | AB | BB | |
| AA | AA_AA | AA_AB | n/a |
| AB | AB_AA | AB_AB | AB_BB |
| BB | n/a | BB_AB | BB_BB |
The allelic effect is examined through a subgroup analysis using only those mother-offspring pairs that had compatible genotypes, i.e. ‘AA_AA’, ‘BB_BB’, and ‘AB_AB’. In doing so, the MFG incompatibility effect is not present and any significance is attributable to the allelic effect.
Step 2. Testing of MFG incompatibility
Step 2 tests the MFG incompatibility effect following each scenario determined in Step 1. A total of 4 scenarios were adopted accordingly to test the MFG incompatibility, see Table 2 for details of these scenarios and the parameterization of the MFG test. SNPs with a small frequency allele (allele frequency p ≤ 0.1) are not tested for the MFG incompatibility effect.
Scenario 1. No allelic effect: f(AA_AA) = f(AB_AB) = f(BB_BB), where function f represents the composite genotype effect.
A logistic regression model was then fitted to all mother-offspring pairs to test the association between the disease and the MFG incompatibility (1 for incompatible pairs and 0 for compatible) adjusting for environmental covariates.
Scenario 2. Allele A effect: f (AA_AA) ≠ f (AB_AB) = f (BB_BB), where the heterozygous genotype has the same effect as one homozygous but significantly different from the other.
To test the MFG incompatibility effect, a logistic regression model was fitted to all mother-offspring pairs adjusting for the individual ‘AA’ genotype effect βAA of either mother or offspring genotype and the father’s allele A contribution λA conditioning on the incompatible MFG.
Scenario 3. Heterozygosity effect: f(AA_AA) = f(BB_BB) ≠ f(AB_AB), where the two homozygous genotypes had the same effect but significantly different from the heterozygous.
To test the MFG incompatibility effect, a logistic regression model was fitted to all mother-offspring pairs adjusting for mother’s ‘AB’ genotype effect βAB and father’s allele A contribution λA conditioning on the incompatible MFG.
Scenario 4. Unequal genotype effects (significantly different effects), including the following two cases.
Table 2. Parameterization of logistic regression model testing MFG incompatibility| Scenario 1. f(AA_AA) = f(AB_AB) = f(BB_BB): no allelic effect | |
| Model effect | Incompatible |
| AB → AB | |
| BB → BB | |
| AA → AA | |
| AA → AB | γ |
| AB → AA | γ |
| AB → BB | γ |
| BB → AB | γ |
| Scenario 2. f(AA_AA) ≠ f(AB_AB) = f(BB_BB): allele A effect | |||
| Model effect | Incompatible | Genotype | Father|incompatibility |
| AB → AB & BB→ BB | |||
| AA → AA | βAA | ||
| AA → AB | γ | βAA | |
| AB → AA | γ | βAA | λA |
| AB → BB | γ | ||
| BB → AB | γ | λA | |
| Scenario 3. f(AA_AA) = f(BB_BB) ≠ f(AB_AB): heterozygosity effect | |||
| Model effect | Incompatible | Genotype | Father|incompatibility |
| AA → AA & BB → BB | |||
| AB → AB | βAB | ||
| AA → AB | γ | ||
| AB → AA | γ | βAB | λA |
| AB → BB | γ | βAB | |
| BB → AB | γ | λA | |
4.1 Strong allelic interaction effect: f(AB_AB) > f(AA_AA) > f(BB_BB) or
f(AB_AB) < f(AA_AA) < f(BB_BB), where the heterozygous effect is the largest or smallest.
A logistic regression model was fitted to all mother-offspring pairs to test the MFG incompatibility adjusting for mother’s AB genotype effect βAB, mother’s AA genotype effect βAA, and father’s allele A contribution λA conditioning on the incompatible MFG.
| Scenario 4.1: Strong interactive allele effect f(AB) > f(AA) > f(BB) or f(AB) < f(AA) < f(BB) | |||
| Model effect | Incompatible | genotype | Father|incompatibility. |
| BB → BB | |||
| AB → AB | βAB | ||
| AA → AA | βAA | ||
| AA → AB | γ | βAA | |
| AB → AA | γ | βAB | λA |
| AB → BB | γ | βAB | |
| BB → AB | γ | λA | |
4.2 Additive or multiplicative allelic effect: f(AA_AA) > f(AB_AB) > f(BB_BB), where the heterozygous effect is between the two homozygous effects.
To test the MFG incompatibility effect, a logistic regression model was fitted to all mother-offspring pairs adjusting for mother’s allele A effect βA and father’s allele A contribution λA conditioning on the incompatible MFG.
It is worthwhile to note that for some SNPs, one minor allele may lead to small counts of the homozygous composite genotype of mother-offspring pairs of that allele, such as small counts of ‘AA_AA’ composite genotype when allele A has a small frequency. As a result, only a small number of samples may be observed in the compatible group with double homozygous minor allele. This often leads to non-significant MFG incompatibility in either Scenario 1 or 2 although it is difficult to distinguish between these two scenarios with a small frequency allele.
| Scenario 4.2: Additive or multiplicative allele effect f(AA )>f(AB) > f(BB) or f(AA) <f(AB) < f(BB) | |||
| Model effect | Incompatible | genotype | Father|incompatibility. |
| BB → BB | |||
| AB → AB | βA | ||
| AA → AA | 2βA or βA2 | ||
| AA → AB | γ | 2βA or βA2 | |
| AB → AA | γ | βA | λA |
| AB → BB | γ | βA | |
| BB → AB | γ | λA | |
Simulation Studies
We conducted simulations to study the power of the 2-stage test on the incompatibility for every scenario in Table 2. In each simulation, we generated 1000 subjects, each of which had a binary indicator for disease phenotypes (Y), MFG incompatibility effect (C), allelic effect (A) and father’s allelic contribution effect given incompatibility (F). The genotype data were generated under the assumption of HWE. The logistic regression was applied to detect the incompatibility effect at the level of 0.05.
Data Generation
We assumed the allele frequency p for allele A and q=1-p for allele B. Table 3 provides the distribution of the MFG under the HWE. In all scenarios, p was generated from a uniform distribution U01. The incompatibility index C was then generated from a Bernoulli distribution with C = 1 for incompatible MFG:
P(C=1) = p2q + pq2 + p2q + pq2 = 2pq,
P(C=0) = 1–2pq = p2 + q2
Other variables were generated according to the scenarios as follows.
No allelic effect. Two probabilities r1 and r2 for having disease with incompatible and compatible MFG, respectively, were generated from uniform distribution U01, and 1000 subjects were generated with probabilities Pr(Y=1|C=1) = r1, Pr(Y =1|C=0) =r2.
Allele A effect. First, six probabilities r1, r2……r6 ~ U 01 of having the disease were generated according to different values of C, A, and F, where F was generated with Bernoulli probabilities Pr(F=1|C=1)=(p2q+pq2)/2pq = 0.5 and Pr(F=1|C=0)=0. A was the ‘AA’ genotype effect of the mother or the fetus or both, and was simulated by Pr(A=1|F=1,C=1)=p, Pr(A=1|F=0, C=1)=p, and Pr(A=1|F=0,C=0)=p3/ (p2+q2). Y was the disease phenotype and was simulated with the probabilities r1, …, r6 as listed in Table 4.
Heterozygosity effect. First, six probabilities r1, r2……r6 of having the disease were generated from uniform distribution U01 according to different values of C, A, F. F was generated by Bernoulli trials with probabilities Pr(F=1|C=1)= (p2q+pq2)/2pq=0.5 and Pr(F=1|C=0)= 0. A was the effect of genotype ‘AB’ in mother or fetus or both and was generated with Pr(A=1|F=1,C=1)=p, Pr(A=1|F=0,C=1)=q, and Pr(A=1|F=0,C=0) =pq/(p2+q2). Y was generated by Bernoulli trials with probabilities r1, …, r6 as listed in Table 4.
Strong allelic interaction effect. First, seven probabilities r1, r2……r7 of having the disease were generated from uniform distribution U01 according to different values of C, A, F to accommodate three genotype effects βAB, βAA and βBB. F was generated with the probabilities Pr(F=1|C=0)=0 and Pr(F=1|C=1) = (p2q+pq2)/2pq = 0.5. A was the additive effect of allele A and was generated with Bernoulli probabilities
Pr (A=1 | F=1, C=1) = p,
Pr (A=0 | F=1, C=1) = 0, Pr (A=-1 | F=1, C=1) = q,
Pr(A=1 | F=0, C=1) = q,
Pr (A=0 | F=0, C=1) = p, Pr (A=-1 | F=0, C=1) = 0,
Pr(A=1 | F=0, C=0) = pq/(p2+q2),
Pr (A=0 | F=0, C=0) = p3/(p2+q2),
Pr (A=-1 | F=0, C=0) = q3/(p2+q2),
where A = 1 for genotype ‘AB’ in mother or fetus, 0 for genotype ‘AA’, and -1 for genotype ‘BB’. Y was generated by Bernoulli trials with probabilities r1,……r7 as in Table 4.
Table 3. Distribution of maternal-fetal genotype under HWE with allele frequencies (p,q).| Mother genotype | Offspring genotype | ||
| AA | AB | BB | |
| AA | p3 | p2q | 0 |
| AB | p2q | pq | pq2 |
| BB | 0 | pq2 | q3 |
| Pr(Y=1) | C | A | F |
| Pr(Y=1) | C | A | F |
| Models in scenarios 2 and 3. | |||
| r1 | 1 | 1 | 1 |
| r2 | 1 | 1 | 0 |
| r3 | 1 | 0 | 1 |
| r4 | 1 | 0 | 0 |
| r5 | 0 | 1 | 1 |
| r6 | 0 | 1 | 0 |
| Models in scenario 4. | |||
| r1 | 1 | 1 | 1 |
| r2 | 1 | 1 | 0 |
| r3 | 1 | -1 | 1 |
| r4 | 1 | 0 | 0 |
| r5 | 0 | 1 | 0 |
| r6 | 0 | 0 | 0 |
| r7 | 0 | 1 | 0 |
Simulation Result
We fitted logistic regression model to the generated data with 1000 subjects and calculated the power of the test defined as out of the total number of simulations the number of times when the associated SNP was selected by statistical significance of the test. In scenario 1, the simulation was conducted with different odds ratio (OR) of disease between compatible and incompatible groups. The power of the study was calculated for each of the OR value 3, 2 , 1, 1/2 or 1/3. The simulation was repeated 5,000 times for each value of OR. In scenarios 2-4, because of the random selection of the allelic effect and father effect, the OR was not controlled at first, but rather calculated after the allelic effect and father effect were generated. The ORs were categorized into different intervals of (0,1/3), 1/31/2) [1/22) [23) and [3∞. The simulation was repeated 50,000 times for each of these scenarios to accommodate the unspecified OR value. The power of the study was calculated for each OR interval. It was observed that the power of test increased with the OR as shown in Table 5 and Figure 1.
Table 5. Power of MFG incompatibility test by OR in Scenarios 2-4 simulatio Odds Ratio (OR)| Scenario | (0,1/3)b | [1/3,1/2)b | [1/2.2)b | [2,3)b | [3,+∞)b | |
| 1 | 99.88% | 96.16% | 0.84% | 97.6% | 99.98% | |
| (4994/5000) | (4808/5000) | (42/5000) | (4880/5000) | (4999/5000) | ||
| 2 | 94.62% | 61.84% | 27.67% | 60.61% | 91.19% | |
| (2234/2361) | (6112/9884) | (6747/24384) | (5702/9408) | (3614/3963) | ||
| 3 | 88.89% | 59.46% | 27.71% | 58.72% | 86.98% | |
| (1665/1873) | (5574/9374) | (7384/26650) | (5452/9284) | (2452/2819) | ||
| 4 | 91.17% | 61.25% | 27.63% | 60.40% | 89.11% | |
| (1486/1621) | (5565/9085) | (7661/27727) | (5516/9133) | (2169/2434) | ||
Figure 1.The power of the two-stage MFG test by the OR between the compatible and incompatible groups for scenarios 1 - 4.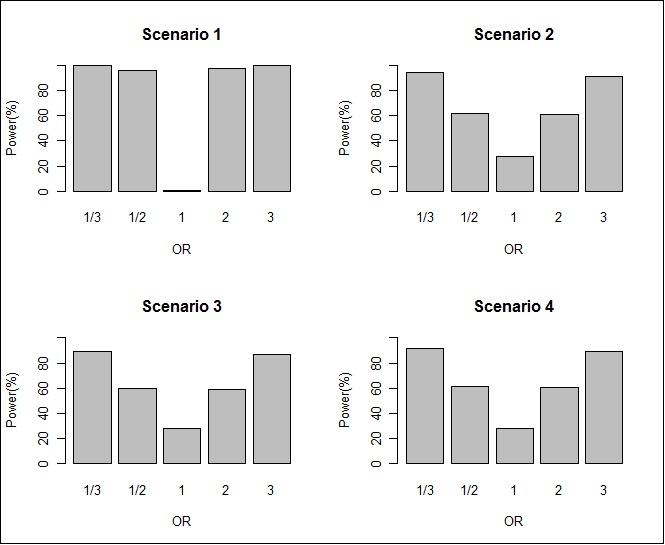
Further simulation was conducted to evaluate the specificity of the test. Ten SNPs were generated and the disease phenotype was generated only by SNP1 in Scenario 1, but independent of SNPs 2-9. The logistic regression model was fitted with 5000 repeats for each OR value, and the power was calculated for each SNP. The simulations demonstrated that the power increased with the OR and that the false positive rate was about 1% (Table 6 and Figure 2).
Table 6. Power of MFG incompatibility test in Scenario 1 simulation based on 5000 repeats with 1000 subjects| SNP | OR=3 | OR = 2 | OR=1.5 | OR=1 |
| SNP1* | 4999 (99.98%) | 4880 (97.60%) | 2881 (57.62%) | 42 (0.84%) |
| SNP2 | 59 (1.18%) | 48 (0.96%) | 41 (0.82%) | 48 (0.96%) |
| SNP3 | 53 (1.06%) | 66 (1.32%) | 39 (0.78%) | 44 (0.88%) |
| SNP4 | 43 (0.86%) | 39 (0.78%) | 52 (1.04%) | 43 (1.04%) |
| SNP5 | 41 (0.82%) | 50 (1.00%) | 43 (0.86%) | 40 (0.80%) |
| SNP6 | 60 (1.20%) | 56 (1.12%) | 40 (0.80%) | 49 (0.98%) |
| SNP7 | 60 (1.20%) | 45 (0.90%) | 44 (0.88%) | 44 (0.88%) |
| SNP8 | 49 (0.98%) | 53 (1.06%) | 53 (1.00%) | 51 (1.02%) |
| SNP9 | 50 (1.00%) | 34 (0.68%) | 38 (0.76%) | 51 (1.02%) |
| SNP10 | 49 (0.98%) | 44 (0.88%) | 44 (0.88%) | 53 (1.06%) |
Figure 2.The specificity of the two-stage method for MFG test by OR.
Application to a Case – Control Study of SGA
We demonstrate our method by studying the MFG incompatibility effect of SNPs on low birth weight in a case-control study of SGA neonates with candidate genes. While the definition of SGA can be found in the literature (Alkalay et al., 1998, Saenger et al., 2007)2,32,, the etiology remains complex. While it was suggested that most of the genetic influence originated in the fetal genome, other results (Ounsted et al. 1988)28 also suggested that not only fetal genes but also genes regulating the maternal uterine environment could be important in determining the size at birth, especially for the SGA neonates.
Insulin-like growth factor (IGF) genes are known to have a major influence on fetal and postnatal growth, and their expression presents in all fetal tissues along the gestation process. On one hand, the cord concentrations of IGF1 and IGF2 have been shown to be highly correlated with birth weight (Ong et al., 2002)27. On the other hand, while some polymorphisms in genes encoding IGF1, IGF2 and their respective receptors have been reported to be associated with birth weight (Vaessen et al. 2002)39, but failed to be confirmed in other studies (Frayling et al. 2002)11. Similarly, the reported association in studies with offspring population remains unconfirmed by one another. So far, no study has combined maternal and fetal genotypes together to study the association between SGA and the incompatibility of the mother and offspring genotypes. In this study, we choose the candidate genes (IGF1 and IGF2 and their receptors) to test the MFG incompatibility effect at each single SNP locus.
Results
Two-stage MFG incompatibility test
In the logistic regression analysis, mother’s age (age), body mass index (BMI) and baby’s sex (sex), were included as environmental covariates as suggested in the literature. Among the 16 SNPs considered for MFG incompatibility analysis, one SNP located at downstream 14463 intron21b of IGF2R gene was found to have a significant allele A effect (Table 7), where the composite genotype ‘AA_AA’ had a significantly different effect from ‘AG_AG’ and ‘GG_GG’ (p = 0.01) after Bonferroni’s multiple-comparison adjustment. We also examined this SNP in mother and offspring populations separately (Table 8) and found that this SNP had a significant genotype effect ‘A/A’ from ‘A/G’ in the mother population as well, but not significant in the offspring population. Further analysis did not reveal any MFG incompatibility effect of this SNP. We further studied the difference between the ‘AA_AA’ and ‘AA_AG’ composite genotypes. The p-value was 0.015, which suggested a significant effect of father’s contribution of allele G given mother genotype ‘AA’ with the Bonferroni’s correction. The remaining 15 SNPs had no significant allelic effect as shown in Table S2Supplementary Material. Among them, one had significant MFG incompatibility effect (Table 9). This SNP is located at downstream 1260 intron2a of IGF1R gene and the incompatible MFG of the SNP decreased the risk for SGA with OR = 0.75 (p=0.033).
Table 7. Test of allelic effect for each SNP| Polymorphism | Compatible Genotype | n* | Testing of Composite Genotype | p-value | ||
| SNP ID | Region | ATG | ||||
| 41410456G/A | IGF2RIntron 21b | 14463 | A/A_A/A | 24 | f(A/A)=f(G/A)=f(G/G) | 0.005 |
| G/A_G/A | 177 | f(A/A)=f(G/A) | 0.015 | |||
| G/G_G/G | 320 | f(A/A)=f(G/G) | 0.008 | |||
| f(G/A)=f(G/G) | 0.644 | |||||
| Polymorphism | Genotype(Samples) | EffectEstimate | OR (95% CI) | p-value | ||
| SNP ID | Region ATG | Overall P-value | ||||
| Mother Population | ||||||
| 42260934A/T | IGF2R | 0.037 | A/A (250) | -0.343 | 0.71 0.51 0.98 | 0.037 |
| Intron17 11258 | T/T (227)A/T (476) | -0.361- | 0.70 0.50 0.97 | 0.033 | ||
| 41410456G/A | IGF2R | 0.037 | A/A (72)G/G (484) | -0.609-0.258 | 0.540.31 0.960.770.58 1.03 | 0.0340.079 |
| Intron21 14463 | G/A (340) | - | ||||
| Offspring Population | ||||||
| 632197358C/A | Intron 1 4560 | 0.040 | A/A ( 0)C/C (51)C/A (851) | N/A-0.580- | 0.56 0.31 1.00 | 0.040 |
| 41410456G/A | IGF2R Intron2114463 | 0.832 | A/A ( 73)G/G (463)A/G (375) | -0.064 0.058- | 0.94 0.55 1.601.06 0.80 1.41 | 0.8160.689 |
| Polymorphism | EffectEstimate | OR (95% CI) | p-value | |
| SNP ID | Region (ATG) | |||
| 40925717 C/T | IGF1R Intron2a (1260) | -0.293 | 0.75 0.570.98 | 0.033 |
Discussion
Recent studies have shown that the MFG incompatibility is associated with several genetic phenotypes, including childhood autism, schizophrenia, etc. Former methods developed for the MFG incompatibility test are based on loglinear regression model on parents-offspring trio genotype data and have been successfully applied to a number of studies. However it is also common that paternal genotypes are unavailable in many studies of only mother-offspring pairs. Therefore, it is desirable to extend the MGF test from parents-offspring trio genotype data to mother-offspring pair genotype data. We have developed a novel two-stage MFG incompatibility test based on logistic regression model that examines the allelic effect of each SNP in the first step and the MFG incompatibility effect in the second step. We also have conducted simulation studies on the power of the test and applied this novel method to a study of the association between the development of SGA and the IGF genes and receptors.
Comparing with the former loglinear MFG incompatibility test, our novel method has the following features. 1). It does not require father’s genotype data, thus is applicable to broad studies where father’s genotype data are unavailable; 2). It tests the allelic effect first and classifies the SNPs into different scenarios by genetic effects of the alleles, which may provide insight on the genetic effects of the SNPs on the phenotypes under investigation; 3). The MFG incompatibility test is conducted based on the allelic effect tested, is more specific for each allele and thus more powerful for each SNP; 4). The power of the MFG incompatibility test increases fast with the OR of the incompatible MFG effect while the false discovery rate is low around 0.01.
Conclusion
This novel two-stage method extends the MFG test from the parents-offspring trio genotype data to mother-offspring pair genotype data and thus makes the MFG test available through an easy-to-understand-easy-to-implement approach to many investigators in a wider range of studies who are not statistical professionals. The examination of the allelic effect may provide clues in the association between the SNP genotype and phenotype of interest, and is easy to interpret.
Supplementary Data:
Supplementary Table 1:
| Clinical and demographic characteristics of subjects by study groupa. | |||
| Control (N=585) | SGA (N=406) | ||
| Maternal age | 25.58 (6.97) | 25.29 (6.04) | |
| Maternal BMI | 24.27 (3.76) | 23.66 (3.99) | |
| Smoking status | |||
| None | 500 (85.47%) | 331 (81.53%) | |
| <1/day | 24 ( 4.10%) | 12 ( 2.96%) | |
| 1-5/day | 54 ( 9.23%) | 53 (13.05%) | |
| 6-10/day | 5 ( 0.85%) | 8 ( 1.97%) | |
| >10/day | 2 ( 0.34%) | 2 ( 0.49%) | |
| Parity (excluding pregnancy in current study)b | |||
| 0 | (44.44%) | 207 (50.99%) | |
| 1 | 184 (31.45%) | 99 (25.38%) | |
| 2 | 90 (15.38%) | 56 (13.79%) | |
| >=3 | 51 ( 8.72%) | 44 (10.84%) | |
| Gestation age (week) | 39.87 (1.12) | 38.45 (2.57) | |
| Birth weight (gram) | 3446.36 (286.27) | 2512.81 (469.07) | |
| Neonatal gender, male % | 51.3 | 46.1 | |
Supplementary Table 2 :
| Polymorphism | Compatible Genotype | n a | Testing of Composite Genotype b | p-value | ||
| SNP ID | Region | ATG | ||||
| 632197358 C/A | IGF1Intron 1 | 4560 | A/A_A/AA/C_A/CC/C_C/C | 025781 | f(C/A)=f(C/C) | 0.10 |
| 40925717 C/T | IGF1RIntron2a | 1260 | C/C_C/CC/T_C/TT/T_T/T | 37815415 | f(C/C)=f(C/T)=f(T/T) | 0.93 |
| 40910576 T/C | IGF1R Intron7b | 5770 | C/C_C/CT/C_T/CT/T_T/T | 16819762 | f(C/C)=f(T/C)=f(T/T) | 0.20 |
| 30729226 A/G | IGF1R Intron8a | 6070 | A/A_A/AA/G_A/GG/G_G/G | 24624544 | f(A/A)=f(A/G)=f(G/G) | 0.52 |
| 40893937 C/T | IGF1R Intron10b | 8050 | C/C_C/CC/T_C/TT/T_T/T | 37316723 | f(C/C)=f(C/T)=f(T/T) | 0.27 |
| 40893931 C/T | IGF1R Exon 11 | 8238 | C/C_C/CC/T_C/TT/T_T/T | 5611155 | f(C/C)=f(C/T)=f(T/T) | 0.12 |
| 44530209 T/C | IGF1R Intron17a | 12443 | C/C_C/CT/C_T/CT/T_T/T | 48189190 | f(C/C)=f(T/C)=f(T/T) | 0.46 |
| 29062007 C/T | IGF2Intron 1 | -177 | C/C_C/CC/T_C/TT/T_T/T | 6001150 | f(C/C)=f(C/T) | 0.90 |
| 29061912 T/C | IGF2Intron 2 | 218 | C/C_C/CT/C_T/CT/T_T/T | 29207280 | f(C/C)=f(T/C)=f(T/T) | 0.06 |
| 42252801 G/A | IGF2RIntron 4b | 2803 | A/A_A/AG/A_G/AG/G_G/G | 100192138 | f(A/A)=f(G/A)=f(G/G) | 0.44 |
| 62092629 G/A | IGF2R Exon 12 | 7575 | A/A_A/AG/A_G/A G/G_G/G | 44203262 | f(A/A)=f(G/A)=f(G/G) | 0.25 |
| 42246208 C/T | IGF2R Intron 16 | 10542 | C/C_C/CC/T_C/TT/T_T/T | 133199102 | f(C/C)=f(C/T)=f(T/T) | 0.52 |
| 42260950 G/A | IGF2R Intron 16 | 10828 | A/A_A/AG/A_G/AG/G_G/G | 6110449 | f(G/A)=f(G/G) | 0.38 |
| 42260934 A/T | IGF2RIntron 17 | 11258 | A/A_A/AA/T_A/TT/T_T/T | 12224299 | f(A/A)=f(A/T)=f(T/T) | 0.44 |
| 42254428 A/G | IGF2RExon 40 | 26813 | A/A_A/AA/G_A/GG/G_G/G | 5521276 | f(A/A)=f(A/G) | 0.13 |
Acknowledgements
This work was partly supported by a contract from the NIH/NICHD to Michigan State University, and by NIH/NICHD intramural funding.
References
- 1.Ainsworth H F, Unwin J, Jamison D L, Cordell H J. (2011) Investigation of maternal effects, maternal-fetal interactions and parent-of-origin effects (imprinting), using mothers and their offspring. , Genet Epidemiol 35(1).
- 2.Alkalay A L, Graham Jr JM, Pomerance J J. (1998) Evaluation of neonates born with intrauterine growth retardation: review and practice guidelines. , J Perinatol 18, 142-151.
- 3.Ashton I K, Eischenk I. (1985) Mckenzie IZ: Insulin-like growth factors (IGF) I and II in human foetal plasma and relationship to gestation age and foetal size during midpregnancy. Acta Endocrinol (Copenh). 110-558.
- 4.Baker J, Liu J P, Robertson E J. (1993) Efstratiadis A: Role of insulin-like growth factors in embryonic and postnatal growth. Cell. 75-73.
- 5.Bonapace G, Concolino D, Formicola S. (2003) Strisciuglio P: A novel mutation in a patient with insulin-like growth factor 1 (IGF-1) deficiency. , J Med Genet 40, 913-917.
- 6.J Lin Chen, Hochner D, H.Semiparametric Maximum Likelihood Methods for Analyzing Genetic and Environmental Effects with Case-Control Mother–Child Pair Data. , Biometrics 68, 869-877.
- 7.Chen J, Zheng H, Wilson M. (2009) Likelihood ratio tests for maternal and fetal genetic effects on obstetric complications. , Genet. Epidemiology 33, 526-538.
- 8.Ylisaukko-oja T Childs, Turunen J A, Rehnström K, Peltonen L, Lange K et al. (2008) Sinsheimer JS: MFG Maximized: Testing for disease-related maternal-fetal genotype incompatibilities using the software package mendel. Abstract at Amer Soc. , Hum. Genet. Conf. Philadelphia
- 9.Cui Y W Fu, K L Sun, Romero R, Wu R. (2007) Mapping nucleotide sequences that encode complex binary disease traits with HapMap. Current Genomics. 8, 307-322.
- 11.Frayling T M, Hattersley A T, McCarthy A, Holly J, Mitchell S M et al. (2002) Ben-Shlomo Y: A putative functional polymorphism in the IGF-I gene: association studies with type 2 diabetes, adult height, glucose tolerance, and fetal growth in U.K. populations. Diabetes. 51, 2313-2316.
- 12.Freemark M. (2006) Regulation of Maternal Metabolism by Pituitary and Placental Hormones: Roles. in Fetal Development and Metabolic Programming. Horm Res 65, 41-49.
- 13.Gluckman P D, Johnson-Barrett J J, Butler J H, Edgar B W, Gunn. (1983) TR: Studies of the insulin-like growth factor I and II by specific radioligands in umbilical cord blood. Clin Endocrinol (Oxf). 19, 405-413.
- 14.G Romero Tromp, Olson R, Lu J M, Xu Q, Parimi Z et al. (2007) Kuivaniemi H: Candidate-gene association study of mothers with pre-eclampsia, and their infants, analyzing 775 SNPs in 190 genes. Human Heredity. 63, 1-16.
- 15.T, John A. (1999) The fetal insulin hypothesis: an alternative explanation of the association of low birth weight with diabetes and cascular disease. Lancet. 353-1789.
- 16.Hayati A R, Cheah F C, Yong J F, Tan A E, Norizah. (2004) WM: The role of serum insulin-like growth factor I (IGF-I) in neonatal outcome. , J. Clin. Pathol 57, 1299-1301.
- 17.Kraft P, Palmer C G, Woodward A J, Turunen J A, Minassian S et al.Sinsheimer JS: RHD maternal-fetal genotype incompatibility and schizophrenia: extending the MFG test to include multiple siblings and birth order. , Eur J Hum Genet 2004, 192-198.
- 18.M S Kramer. (1987) Determinants of low birth weight: methodological assessment and meta-analysis. , Bull World health Org 65-663.
- 19.Kulich V, Kout M. (1967) Hemolytic disease of a newborn caused by anti-k antibody. Cesk. Pediatr 22, 823-826.
- 20.Langford K S, Nicolaides K H, Jones J, Abbas A, McGregor A M et al. (1995) Serum insulin-like growth factor-binding protein-3 (IGFBP-3) levels and IGFBP-3 protease activity in normal, abnormal and multiple human pregnancy. , J Clin Endocrinol Metab 80, 21-27.
- 21.Li S, Lu Q, Fu W, Romero R, Cui Y. (2009) A regularized regression approach for dissecting genetic conflicts that increase disease risk in pregnancy. , Stat Appl Genet Mol Biol. 8(1): Article 45.
- 22.Little R E. (1987) Sing CF: Genetic and environmental influences on human birthweight. , Am J Hum Genet 40, 512-526.
- 23.Ludwig T, Eggenschwiler J, Fisher P, D'Ercole A J, Davenport M L. (1996) Efstratiadis A: Mouse mutants lacking the type 2 IGF receptor (IGF2R) are rescued from perinatal lethality. in IGF2 and IGF1R null backgrounds. Dev Biol 177-517.
- 24.Magnus P. (1984) Caused of variation in birth weight: a study of offspring of twins. , Clin Genet 25, 15-24.
- 25.Minassian S L, Palmer C G.Sinsheimer JS: An exact maternal-fetal genotype incompatibility (MFG) test. , Genet Epidemiol 2005, 83-95.
- 26.Minassian S L, Palmer C G, Turunen J A, Paunio T, Lönngvist J et al.Sinsheimer JS: Incorporating serotypes into family based association studies using the MFG test. Ann Hum Genet. 2006, 541-543.
- 27.Ong K, Kratzsch J, Kiess W, Costello M, Scott C et al. (2000) Size at birth and cord blood levels of insulin, insulin-like growth factor I (IGF-I), IGF-II, IGF-binding protein-1 (IGFBP-1), IGFBP-3, and the soluble IGF-II/mannose-6-phosphate receptor in term human infants: The ALSPAC Study Team: Avon Longitudinal Study of Pregnancy and Childhood. , J Clin Endocrinol Metab 85, 4266-426.
- 28.Ounsted M, Scott A. (1988) Moar VA: Constrained and unconstrained fetal growth: associations with some biological and pathological factors. Ann Hum Biol. 15, 119-129.
- 29.Palmer C G, Turunen J A, Sinsheimer J S, Minassian S, Paunio T et al. (2002) JA: RHD maternal-fetal genotype incompatibility increases schizophrenia susceptibility. , Am J Hum Genet 2002, 1312-9.
- 30.Palmer C G S Hsieh, H J Reed, E F Lonnqvist, Peltonen J, Woodward L et al. (2006) . , HLA-B Maternal-Fetal Genotype Matching Increases Risk of Schizophrenia. The American Journal of Human Genetics 79, 710-715.
- 31.Reece E A, Wizniter A, E Homko CJ Le, Behrman H, Spencer E M. (1994) The relation between human fetal growth and fetal blood levels of insulin-like growth factors I and II, their binding proteins, and receptors. Obstet Gynecol. 84-88.
- 32.Saenger P, Czernichow P, Hughes L, Reiter E O. (2007) Small for Gestational Age: Short Stature and Beyond. Endocrine Reviews. 28, 219-251.
- 33.Salet J, Colombani J, Girot R, Beaufils F, Lejeune C et al. (1973) Neonatal thrombopenia with feto maternal incompatibility in the HL-A system. Rev Fr Transfus. 243-50.
- 34.Salet J, Colombani J, Fajgenbaum J, Beaufils F, Lejeune C et al. (1973) Neonatal thrombopenia with feto-maternal incompatibility in the HL-A system. Arch Fr Pediatr. 30, 533-40.
- 35.Shi M, D M Umbach, Vermeulen S, C R Weinberg. (2008) Making the most of case-mother/control-mother studies. , Amer. J. Epidemiology 168, 541-547.
- 36.Sinsheimer J S, Palmer C G, Woodward. (2003) JA: Detecting genotype combinations that increase risk for disease: maternal-fetal genotype incompatibility test. Genet Epidemiol. Jan;24 1-13.
- 37.C, R, Chatterjee L. (2004) Analysis of case-control studies of genetic and environmental factors with missing genetic information and haplotype-phase ambiguity. Genetic Epidemiology. 29, 108-127.
- 38.Stubbs E G, Ritvo E R, Mason-Brothers.A: Autism and shared parental HLA antigens. Am Acad Child Psychiatry. 1985, 182-185.
- 39.Vaessen N, Janssen J A, Heutink P, Hofman A, Lamberts S W et al. (2002) Duijn CM: Association between genetic variation in the gene for insulin-like growth factor-l and low birthweight. Lancet. 359-1036.
- 40.Verhaeghe J, R Van Bree, E Van Herck, Laureys J, Bouillon R et al. (1993) Assche FA: C-peptide, insulin-like growth factors I and II, and insulin-like growth factor binding protein-1 in umbilical cord serum: correlations with birthweight. , Am J Obstet Gynecol 169-89.
Cited by (4)
This article has been cited by 4 scholarly works according to:
Citing Articles:
Yuang Tian, Hong Zhang, Alexandre Bureau, H. Hochner, Jinbo Chen - Journal of Statistical Planning and Inference (2024) Semantic Scholar
Journal of Statistical Planning and Inference (2024) Crossref
JUSTC (2022) Crossref
Yanan Zhao, Weiqi Yang, Hong Zhang - JUSTC (2022) Semantic Scholar
Genetic Epidemiology (2021) Crossref
Kai Zhang, Hong Zhang, H. Hochner, Jinbo Chen - Genetic Epidemiology (2021) Semantic Scholar
Yuehua Cui, Haitao Yang - Physics of Life Reviews (2017) Semantic Scholar
