Abstract
The compound 1,4-dimethylnaphthalene, originally isolated from dormant potatoes, is currently in use as a commercial sprout inhibitor. Growers and processors report a reduction in fungal infections in potatoes treated with DMN resulting in increased yields. To assess the effects of DMN on fungal growth a culture of Fusarium oxysporum was isolated from potato tubers and identified via DNA fingerprinting using the 18ITS ribosomal region. Growth of F. oxysporum was inhibited by 31% after four days of exposure to DMN but overall rate of spore germination was not affected by DMN treatment. The growth of additional fungi, including Alternaria alternata, Aspergillus niger, Epicoccum nigrum, Gnomoniopsis smithogilvyi, Phoma medicaginis, and Pythium ultimum was inhibited by DMN as was suppression of sporulation in A. niger. These results suggest that DMN is fungistatic at the application levels examined.
Author Contributions
Academic Editor: Abubaker Haroun Mohamed Adam, Department of Crop Science (Agronomy), College of Agriculture, Bahri University- Alkadaru- Khartoum -Sudan, Sudan.
Checked for plagiarism: Yes
Review by: Single-blind
Copyright © 2019 Michael Campbell, et al.
 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Competing interests
The authors have declared that no competing interests exist.
Citation:
Introduction
Harvested potatoes are often treated with sprout regulators to prolong storage. The synthetic compound isopropyl-(N-3-chloro-phenyl)-carbamate (Chlorpropham) is widely used to prevent sprouting on stored potatoes. However, the inability to use Chlorpropham on potato tubers used as seed, and concerns about residue levels on tubers used for food, has created an interest in finding alternative compounds11. The naturally occurring volatile carbon compound, 1,4 dimethyl naphthalene (DMN), is an effective sprout control agent when applied to harvested potato tubers 1, 7, 16. Growers who utilize DMN demonstrate reduction of potato sprout growth, prolonged storage, and often see a reduction in product loss due to decrease in fungal pathogens. It is unclear if the reduction of disease in storage is a function of DMN directly on pathogens or if DMN induces a response in tubers that increases pathogen resistance. Gene expression analysis on potato tubers treated with DMN did reveal an increase in genes associated with pathogen response 3 but the interaction of DMN directly with fungal pathogens has been poorly assessed.
Essential oils derived from plant material, in many instances containing a complex mixture of hydrophobic molecules, have been shown to have antimicrobial and antifungal activity 2, 5, 12, 18. DMN is a polycyclic aromatic hydrocarbon (PAH) and hydrophobic molecule that is found in some essential oil extracts. Given that growers see a reduction in loss due to disease following treatment with DMN, and that essential oils have been shown to have antimicrobial activity, we undertook an evaluation of fungal growth in response to DMN exposure.
Materials and Methods
Plant Material and Isolation of Fusarium Cultures
Potato tubers (Solanum tuberosum cv. La Chipper) were harvested in the fall of 2016 in Erie County, Pennsylvania. Tubers were allowed to undergo skin for set for 14 days at room temperature and then stored at 8 ºC for five months. Skin scrapings were placed onto Nash/Synder agar plates for selection of Fusarium species10. Following two days of growth at 22 ºC putative Fusarium isolates were plated onto potato dextrose agar (PDA). Isolated cultures were grown at 22 ºC and total DNA was isolated using a PowerSoil DNA Isolation Kit (www.Miobio.com). DNA was quantified using as spectrophotometer and the D1/D2 region of the 26S rRNA gene was amplified using the primers NL1 (GCATATCAATAAGCGGAGGA) and NL4 (GGTCCGTGTTTCAAGACGG 8. An additional genomic region encoding for the 18 ITS was amplified using the primers ITS1 (CTTGGTCATTTAGAGGAAGTAA) and ITS2 (GCGTTCTTCATCGATGC). PCR products were purified and then sent to Retrogen (http://www.retrogen.com) for Sanger sequencing using the same primers used for PCR amplification. DNA sequences were then analyzed using BLASTn against the NCBI database using high similarity and a search restricted to fungi.
Additional Fungal Strains
The fungal strains Aspergillus niger (ATCC: 16888) and Pythium ultimum(ATCC: 58811) were purchased from American Type Culture Collection (www.atcc.org) and maintained on potato dextrose agar (PDA). Alternaria alternatawas isolated from leaves of hardy kiwi (Actinidia arguta) grown at the Lake Erie Regional Grape Research and Extension Center (LERGREC) located in North East, PA. Hardy kiwi leaves were harvested and placed in closed petri dishes until fungal hyphae were visible. Hyphae were then scraped on to PDA and single isolates were selected and maintained on PDA. Epicoccum nigrum, Gnomoniopsis smithogilvyi, and Phoma medicaginis were isolated from phloem tissue obtained from American Chestnut trees located at (LERGREC) that exhibited cankers. Cultures were maintained on PDA and isolates were identified using DNA barcodes based on the 18ITS ribosomal region. Single culture isolates were grown on PDA, and DNA was isolated using a PowerSoil DNA Isolation Kit. DNA was quantified using as spectrophotometer and PCR was accomplished using 18 ITS primers described above. Sanger sequencing was conducted on the purified PCR products and results were evaluated using BLASTn against the NBCI database narrowed to fungal accessions.
Treatment of Fungal Strains with DMN
Fungal cultures were plated onto PDA plates and placed in 9.5 Liter BBL GasPak chamber (http://www.bd.com/ds/productCenter/ES-Gaspak.asp) containing filter paper strips that had 22.5 ul of 1,4-dimethylnaphthalene spotted on to it. Control chambers contained 22.5 ul of water in place of DMN. The chambers were placed at 22 ºC and incubated for two days. The rate of application of DMN was selected based on previous experiments with potatoes that have shown that this application rate results in a DMN residue level on potatoes similar to that found in commercial storage units3.
Fungal spores from Fusarium oxysporum and Aspergillus niger were isolated using a modification of the procedure of Espinel-Ingroff and Kerkering 4. A sporulating culture was washed with 1 ml of sterile water. The water was transferred to a microfuge tube, centrifuged at 9400 g for 1 min and the supernatant was transferred to a sterile microfuge tube. A 500 ul sample was placed into a sterile spectrophotometry cuvette and percent transmittance at 530 nm was determined and then adjusted to 75 ± 1%. The solution was then diluted 1:100 with sterile water and 10 ul was plated on PDA agar and the cultures were incubated at 25 ºC for 24 hrs with and without DMN. Germination was determined by the presence of mycelia and amount of growth was determined by measuring size of the mycelial spread for each germination.
Cultures of A. alternata, A. niger, E. nigrum, G. smithogilvyi, P. medicaginis, and P. ultimum were washed with 1 to 3 ml of sterile water and liquid was placed in a sterile tube and diluted 1:100 with water. Following dilution, 100 ul sample was spread on to PDA and then treated in a GasPak chamber as described for Fusarium oxysporum. Cultures were photographed after two days of exposure to DMN or the water control.
Cultures of Alternaria alternata were allowed to grow across the surface of a plate containing PDA. Using a sterile scalpel mycelial mats of about 0.5 cm were placed on to fresh PDA plates. One set of plates was transferred to a to GasPak chamber for DMN for 24 hrs and another segment of the same culture was placed in a GasPak chamber and exposed to water. Sporulation was assessed after 24 hrs.
Results and Discussion
Isolation of Fusarium Species from Potato and Growth in Response to DMN
Species identification of the isolated strain of Fusarium was based on DNA sequence of the ITS and NL amplified regions. The ITS region revealed similarity to Fusarium oxysporum while the NL regions demonstrated sequence similarity to F. oxysporum sp. lycopersici (Table 1).
Table 1. Blast N results for PCR products generated from DNA isolated from fungal cultures. Descriptions are from the NCBI database accession associated with specific E values.| Species | Primer Type | NBCI | E value | Accession |
|---|---|---|---|---|
| Alternaria alternata | 18 ITS Forward | Alternaria alternata isolate UFSMLB09 internal transcribed spacer 1, partial sequence; 5.8S ribosomal RNA gene and internal transcribed spacer 2, complete sequence; and large subunit ribosomal RNA gene, partial sequence | 5.00E-107 | MH819177.1 |
| 18 ITS Reverse | Alternaria alternata isolate ALT2 small subunit ribosomal RNA gene, partial sequence; internal transcribed spacer 1, 5.8S ribosomal RNA gene, and internal transcribed spacer 2, complete sequence; and large subunit ribosomal RNA gene, partial sequence | 4.00E-113 | MH578598.1 | |
| Epicoccum nigrum | 18 ITS Forward | Epicoccum nigrum strain CCTU118 internal transcribed spacer 1, partial sequence; 5.8S ribosomal RNA gene, complete sequence; and internal transcribed spacer 2, partial sequence | 3.00E-97 | MH_758719.1 |
| 18 ITS Reverse | Epicoccum nigrum isolate Au-V-1.1 small subunit ribosomal RNA gene, partial sequence; internal transcribed spacer 1 and 5.8S ribosomal RNA gene, complete sequence; and internal transcribed spacer 2, partial sequence | 1.00E-96 | MF_475942.1 | |
| Fusarium oxysporum | ITFS Forward | Fusarium oxysporum small subunit ribosomal RNA gene, partial sequence; internal transcribed spacer 1 and 5.8S ribosomal RNA gene, complete sequence; and internal transcribed spacer 2, partial sequence | 2.00E-101 | MG252282.1 |
| ITFS Reverse | Fusarium oxysporum isolate DSM 106834 small subunit ribosomal RNA gene, partial sequence; internal transcribed spacer 1, 5.8S ribosomal RNA gene, and internal transcribed spacer 2, complete sequence; and large subunit ribosomal RNA gene, partial sequence | 2.00E-106 | MH575293.1 | |
| NL1 | Fusarium oxysporum f. sp. lycopersici strain CBS 758.68 large subunit ribosomal RNA gene, partial sequence | 0.00E+00 | MH878407.1 | |
| NL2 | Fusarium oxysporum f. sp. lycopersici strain CBS 130307 large subunit ribosomal RNA gene, partial sequence | 0.00E+00 | MH877312.1 | |
| Gnomoniopsis smithogilvyi | 18 ITS Forward | Gnomoniopsis smithogilvyi isolate THDAS 2016_1120A 18S ribosomal RNA gene, partial sequence; internal transcribed spacer 1, 5.8S ribosomal RNA gene, and internal transcribed spacer 2, complete sequence; and 28S ribosomal RNA gene, partial sequence | 9.00E-119 | KY_695232.1 |
| 18 ITS Reverse | Gnomoniopsis smithogilvyi culture-collection ICMP:14040 18S ribosomal RNA gene, partial sequence; internal transcribed spacer 1, 5.8S ribosomal RNA gene, and internal transcribed spacer 2, complete sequence; and 28S ribosomal RNA gene, partial sequence | 1.00E-126 | KC_145868.1 | |
| Phoma medicaginis | 18 ITS Forward | Phoma medicaginis strain 7_ITS1F_E06_17Q 18S ribosomal RNA gene, partial sequence; internal transcribed spacer 1, 5.8S ribosomal RNA gene, and internal transcribed spacer 2, complete sequence; and 28S ribosomal RNA gene, partial sequence | 5.00E-100 | KF_293990.1 |
| 18 ITS Reverse | Phoma medicaginis strain 7_ITS4_E05_17Q 18S ribosomal RNA gene, partial sequence; internal transcribed spacer 1 and 5.8S ribosomal RNA gene, complete sequence; and internal transcribed spacer 2, partial sequence | 2.00E-103 | KF_293988.1 |
The Fusarium oxysporum isolate, when grown in the presence of DMN for four days, exhibited a decrease in growth (Figure 1). The average size for Fusarium mycelia in the controls was 9.8 ± 0.7 cm while exposure to DMN resulted in an average growth size to 3.0 ± 0.5 cm. The difference was statistically significant based on T-test analysis. However, the difference in the number of spores that germinated in the presence of DMN, compared to the controls, was not statistically significant. Suggesting that, at the concentration used here, DMN is not fungicidal but fungistatic. DMN did not kill the Fusarium cultures but inhibited mycelial growth rates. The ability of DMN to suppress Fusarium growth is significant because Fusarium spp. are the causative agents of rots and wilts in potato15. While no PAH has been shown to specifically inhibit the growth of Fusarium, mixtures of hydrophobic oils from clove and lemon grass have been shown to decrease the growth of F. oxysporum sp. Lycopersici 17.
Figure 1.Fusarium oxysporum isolated from potato tubers grown the presence of DMN for two days.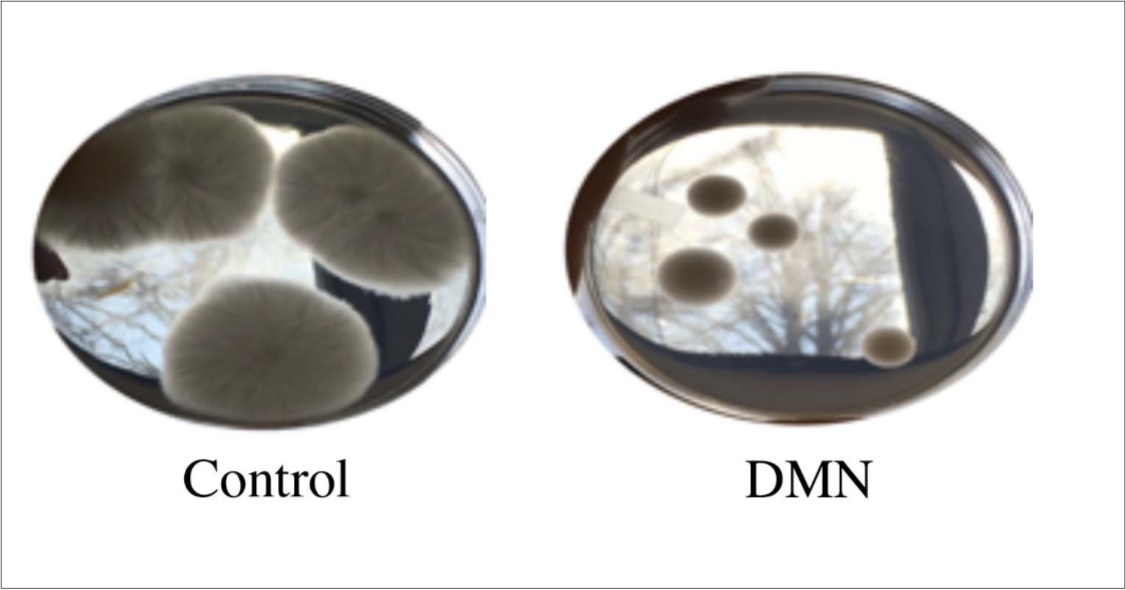
DMN Effects of Multiple Fungal Species
Treatment of Alternaria alternata, Aspergillus niger, E. nigrum, G. smithogilvyi, P. medicaginis, and P. ultimum with DMN resulted in suppression of mycelial growth (Figure 2). The morphology of the mycelium prevents quantitative assessment of the growth rates. The mycelial growth after DMN exposure was less than 5 mm for A. alternata,E. nigrum, and P. medicaginis. The diversity of the species examined ranged from the Ascomycota (A. alternaria, A. niger, E. nigrum, G. smithogilvyi, and P. medicaginis) to the oomycota (P. ultimum) suggesting that DMN suppression of fungal growth appears to be a broad-spectrum response and does not involve inhibition of species-specific pathways. Similar fungistatic activity was found with mixtures of essential 6, 9, 13, 14 Complex mixtures of essential oils may contain polycyclic hydrocarbons (PAHs) with structure similarity to DMN but there is limited literature surrounding substituted naphthalene and mycelial inhibition. Investigations have been conducted using a number of fungal species to mitigate PAH soil contaminants via metabolism but the effect of PAH on rate of growth of fungi was not evaluated.
Figure 2.Fungal cultures grown in the presence of DMN for two days: a. Alternaria alternata, b. Aspergillus niger (ATCC: 16888), c. Epicoccum nigrum, d. Gnomoniopsis smithogilvyi, e. Phoma medicaginis, f. Pythium ultimum (ATCC: 58811)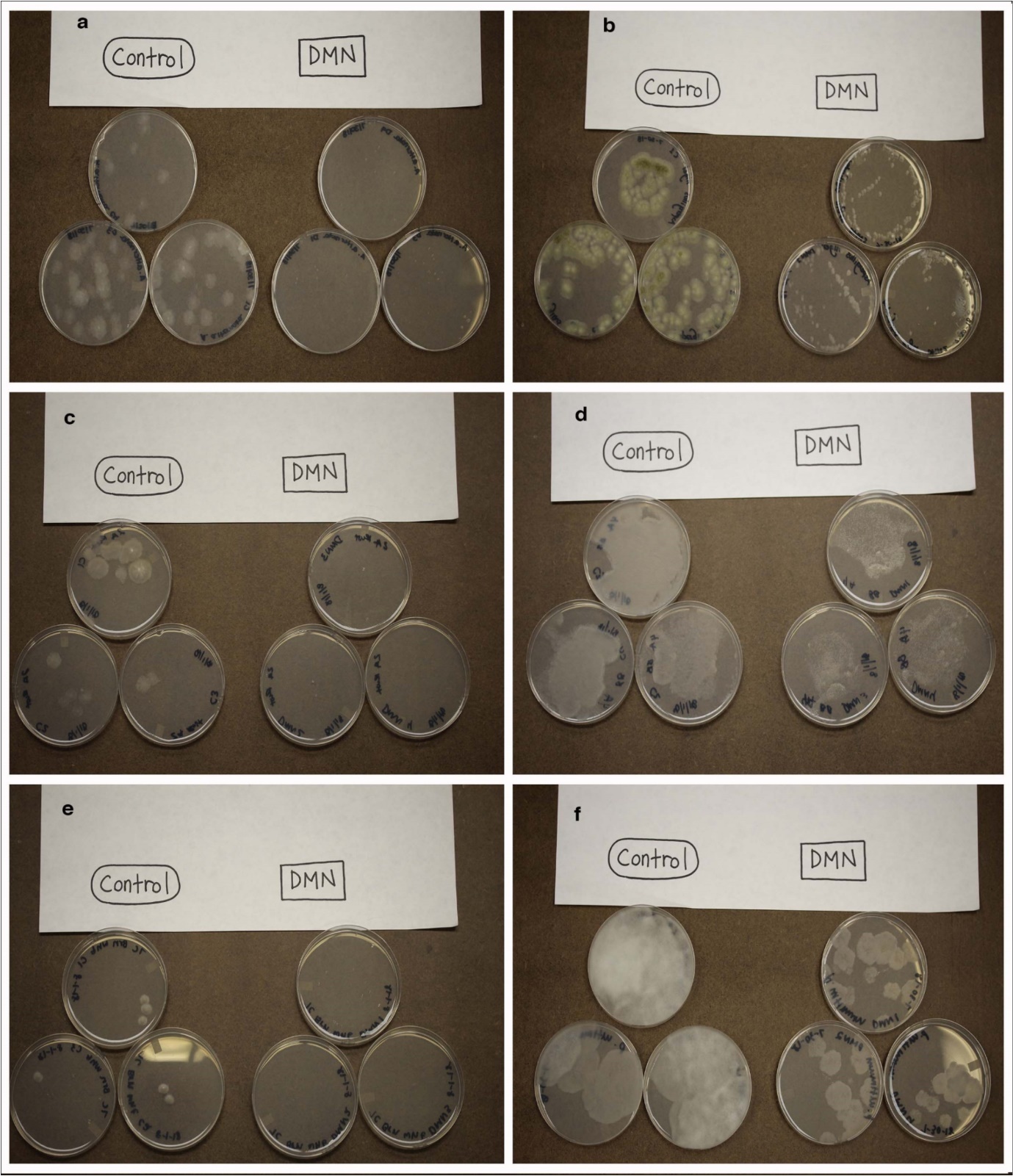
DMN and Sporulation
Treatment of A. Alternaria with DMN resulted in a decrease in sporulation compared to controls (Figure 3). The suppression of sporulation may be a function of reduced growth rate by DMN and not a result in the alteration of the sporulation developmental pathway directly.
Figure 3.Mature of cultures of Aspergillus niger (ATCC: 16888) grown in the presence of DMN showing suppression of sporulation.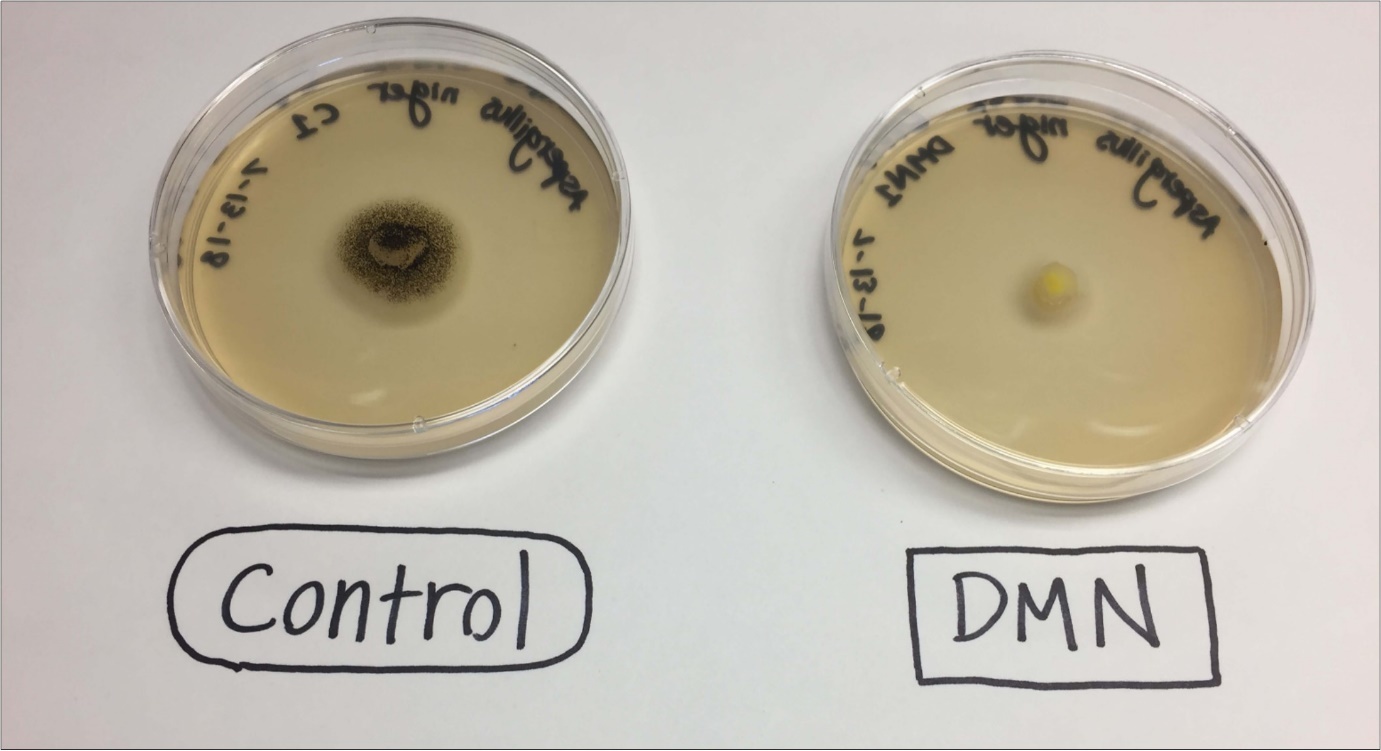
Conclusion
DMN, a compound that is utilized as a sprout regulator in the potato industry, has fungistatic activity against species in the Ascomycota and Oomycota. At the dosage utilized in the potato industry to prevent sprouting, DMN does not inhibit germination of fungal spores nor kill fungal cultures. However, DMN inhibited mycelial growth of all fungal species tested and decreased the rate of sporulation in A. alternaria. Thus, anecdotal evidence where growers and processors find a reduction in fungal disease in potato sheds is supported by the fungistatic effects of DMN.
Acknowledgments
The work was supported in part by grants from the 1,4-Group of Meridian, ID and by the Penn State Behrend Undergraduate Research program.
References
- 1.Beveridge J, Dalziel J, Duncan H. (1981) Dimethylnaphthalene as a sprout suppressent for seed and ware potatoes. , Potato Res 24, 77-88.
- 2.J R Calo, P G Crandall, C A O'Bryan, S C Ricke. (2015) Essential oils as antimicrobials in food systems – A review. , Food Control 54, 111-119.
- 3.M A Campbell, D’Annibale O. (2016) Exposure of Potato Tuber to Varying Concentrations of 1,4-Dimethylnaphthalene Decrease the Expression of Transcripts for Plastid Proteins. , American Journal of Potato Research 93(3), 278-287.
- 4.Espinel-Ingroff A, Kerkering T M. (1991) Spectrophotometric method of inoculum preparation for the in vitro susceptibility testing of filamentous fungi. , Journal of Clinical Microbiology 29(2), 393-394.
- 5.Hammer K A, Carson C F, Riley T V. (1999) Antimicrobial activity of essential oils and other plant extracts. , Journal of Applied Microbiology 86(6), 985-990.
- 6.Khaledi N, Taheri P, Tarighi S. (2015) Antifungal activity of various essential oils against Rhizoctonia solani and Macrophomina phaseolina as major bean pathogens. , Journal of Applied Microbiology 118(3), 704-717.
- 7.G E Kleinkopf, N A Oberg, N L Olsen. (2003) Sprout inhibition in storage: Current status, new chemistries, and natural compounds. , American J. potato res 80(5), 317-327.
- 8.Lopandic K, Zelger S, L K Bánszky, Eliskases-Lechner F, Prillinger H. (2006) Identification of yeasts associated with milk products using traditional and molecular techniques." Food microbiology. 23(4), 341-350.
- 9.Mimica-Dukić N, Božin B, Soković M, Mihajlović B, Matavulj M. (2003) Antimicrobial and Antioxidant Activities of Three Mentha Species Essential Oils. , Planta Med 69(05), 413-419.
- 10.S M Nash, W C Snyder. (1962) Quantitative estimations by plate counts of propagules of the Bean root rot Fusarium in field soils. , Phytopathology 52(6), 567-572.
- 11.Paul V, Ezekiel R, Pandey R. (2016) Sprout suppression on potato: need to look beyond CIPC for more effective and safer alternatives. , Journal of Food Science and Technology 53(1), 1-18.
- 12.Prakash B, Kedia A, P K Mishra, N K Dubey. (2015) Plant essential oils as food preservatives to control moulds, mycotoxin contamination and oxidative deterioration of agri-food commodities – Potentials and challenges. , Food Control 47, 381-391.
- 13.M S Rakotonirainy, Lavédrine B. (2005) Screening for antifungal activity of essential oils and related compounds to control the biocontamination in libraries and archives storage areas. , International Biodeterioration & Biodegradation 55(2), 141-147.
- 14.Reuveni A F, Putievsky R E. (1984) Fungistatic Activity of Essential Oils from Ocimum basilicum Chemotypes. , Journal of Phytopathology 110(1), 20-22.
- 16.Knowles R, Knowles N, Haines M M. (2005) 1,4-Dimethylnaphthalene treatment of seed potatoes affects tuber size distribution. , American Journal of Potato Research 82(3), 179-190.
Cited by (5)
- 1.Dobry Emily P., Rutter Michael A., Campbell Michael, 2023, The Fungal Pathogen Gnomoniopsis castaneae Induces Damaging Cankers in Multiple Domestic Fagaceae Species, Phytopathology®, (), 10.1094/PHYTO-12-22-0451-SC
- 2.Dobry Emily, Campbell Michael, 2023, Gnomoniopsis castaneae: An emerging plant pathogen and global threat to chestnut systems, Plant Pathology, 72(2), 218, 10.1111/ppa.13670
- 3.Santos Mirelle Nayana de Sousa, de Araújo Nicolas Oliveira, de Araujo Fernanda Ferreira, da Silva Mariana Aparecida, Barbosa Samuel Lessa, et al, 2023, Sprout-suppressing 1,4-dimethylnaphthalene treatment reduces dry rot infection in potato tubers during postharvest storage, Postharvest Biology and Technology, 205(), 112485, 10.1016/j.postharvbio.2023.112485
- 4.Krause Marcelo Rodrigo, Ferreira de Araujo Fernanda, Ferreira Moreira Karoliny, Oliveira de Araújo Nícolas, Paulo de Jesus Tello Jean, et al, 2023, Carbohydrate metabolism dynamic in chlorpropham- and 1,4-dimethylnaphthalene-treated potatoes and its effect on the browning of French fries, Food Chemistry, 429(), 136718, 10.1016/j.foodchem.2023.136718
- 5.Gumbo Nyasha, Magwaza Lembe Samukelo, Ngobese Nomali Ziphorah, 2021, Evaluating Ecologically Acceptable Sprout Suppressants for Enhancing Dormancy and Potato Storability: A Review, Plants, 10(11), 2307, 10.3390/plants10112307
