Docking Studies of HIV-1 Reverse Transcriptase and HIV-1 Protease with Phytocompounds of Carissa Carandas L.
Abstract
Background
Carissa carandas L. is a well-known wild fruit plant distributed through-out the India and also present in other countries. The fruits are rich in nutrients and minerals. A number of medicinally important phytochemicals such as carrisone, carindone, carandinol, lupeol, scopoletin, stigmasterol, β-sitosterol, myo-inositol, β-amyrin, Des-n-methylnoracronycine etc. have been reported from the extract of this plant. Being safe and cost effective molecules, the activity of phytochemicals against HIV-1 enzymes needs to be screened.
Objective
The aim of this study was to screen the potent phytocompound of C. carandas against human immunodeficiency virus-1 using docking method.
Methods
Total nine compounds viz. carandinol, caridone, carrisone, lupeol, p-coumaric acid, gallic acid, rutin, scopoletin and ursolic acid were used for in-silico study towards drug development against human immunodeficiency virus-1 reverse transcriptase (HIV-1RT; PDB ID: 1REV) and human immunodeficiency virus-1 protease (PDB ID:1EBY) using Autodock software.
Results
The qualitative characterization of the extracts showed the presence of a number of phytochemicals such as phenolics, flavonoids, alkaloids, terepnoids, terpenes, steroids, glycosides etc. Carandinol was observed as most effective anti-HIV-1 molecule having lowest binding energy and small inhibition coefficient. Another compound, p-coumaric acid, showed least effectiveness against human immunodeficiency virus- 1 reverse transcriptase or human immunodeficiency virus-1 protease showing highest binding energy and inhibition coefficients among all the evaluated phytocompounds.
Conclusion
The in-silico study demonstrated that some phytoconstituents of C. carandas exhibit potential anti-human immunodeficiency virus -1 activity and hence can be optimized to develop as a drug candidate in future.
Author Contributions
Academic Editor: Shivaji Kashinath Jadhav, MapMyGenome India Limited, Hyderabad Previous experience, Sandor Life Sciences Pvt Ltd/ NIMR, Indian Council of Medical Research, Goa, NIRRH ICMR Mumbai, India.
Checked for plagiarism: Yes
Review by: Single-blind
Copyright © 2019 Reetika Singh, et al.
 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Competing interests
The authors have declared that no competing interests exist.
Citation:
Introduction
Human immunodeficiency virus (HIV) is one of the most austere and deadly infectious viruses with disastrous concerns 1, 2. Infections with HIV caused Acquired Immune Deficiency Syndrome (AIDS) and it has been considered as a pandemic disease 1, 3. HIV-1 is more common and high infectious than HIV-2. HIV-1 has the ability to mutate and change their genetic makeup over the time like many other viruses. There are many genetically distinct subgroups derived from the main type of HIV. HIV-1 is predominant type of HIV and 95% of total HIV infectious person infected with HIV-1. HIV-1 and HIV-2 viruses are genetically different from each other by almost 55% 4. HIV-1 has four subtypes placed in different groups such as M, N, O and P. Group M is the most dominant and infectious in nature and this group have several strains such as A,B, C, D, F, G, H, J, K and Circulating Recombinant Forms (CRFs). CRFs are hybrid viruses formed as a result of combination of the genetic materials of two subtypes 5, 6. HIV has infected to about 70 million people all over the world since the beginning of the epidemic and has caused mortality to about 50% of the infected population. Till the end of 2017, about 37 million people were reported to harbour HIV-1 infection 7. About 40 million peoples are suffering from HIV infection and Africa region is a serious victim with 25 million peoples. Every year about 2.4 million deaths occurred due to HIV and AIDS 4. According to the National Aids Control Organisation (NACO, India) 2015 reports, in adults HIV occurrence in India was about 0.26% (0.22%–0.32%). HIV prevalence was about 0.30% in males and 0.22% in females in India. Among the Indian states, Manipur has shown the highest estimated adult HIV prevalence of 1.15% and some states have HIV prevalence below 0.20% 8.
Reverse transcriptase (RT) is most important enzyme for the retroviruses to complete their life cycle 9. Reverse transcriptase (HIV-1 RT) contains 1000 amino acids and is an asymmetric heterodimer. The larger subunit known as p66 contains 560 amino acid long and performing all the enzymatic activities of the RT 10. Two active sites namely polymerase and ribonuclease H have been found on larger subunit (p66). The smaller subunit p51 contains 440 amino acids and it is deliberated to stabilize the heterodimer but this unit also may take part in t-RNA primer binding 10, 11, 12. The HIV-1RT catalyses replication of viral genome (mRNA) converting it into proviral DNA (cDNA) which gets integrated into the human genome present into the nucleus of an infected cell. The integration is catalysed by HIV-1 integrase. The ribonuclease H-domain of the larger subunit (p66) of HIV-1 RT degrades retroviral RNA while synthesising c DNA 10, 12. Most of the anti-HIV-1 drugs have targeted the RT so as to block cDNA synthesis and hence further sequence of reactions concerning retroviral replication.
HIV-1 protease is another key enzyme which is essential for the HIV life-cycle 13. The protease cleaves the newly translated chain and generates required protein components to assemble the infectious HIV virions 13, 14. HIV virions become uninfectious without operative HIV protease 15, 16. Thus, the inhibition of the activity of HIV-1 or the mutation of the active site residues of HIV protease may disrupt the packaging and maturation of the viruses 17. Therefore, HIV protease becomes one of the main targets towards design and development of many new HIV-1 drugs in addition to the reverse transcriptase and integrase enzymes 18.
Various synthetic drugs are in practice against reverse transcriptase such as nucleoside / nucleotide reverse transcriptase inhibitors (Abacavir, Didanosine, Emtricitabine, Lamivudine, Stavudine, Tenofovir, Zidovudine) and non-nucleoside reverse transcriptase inhibitors (NNRTI) (efavirenze, nevirapine, delavirdine and etravirdine). The drugs against HIV-1 protease include saquinavir, ritonavir, indinavir, and nelfinavir. The applications of these synthetic drugs are reported to develop drug resistance in HIV and also the emergence of the new species of the virus 19, 20. The approved anti HIV synthetic drugs are effective in the early phase but during the course of time the efficacy starts decreasing. Therefore, it is a necessity to develop a cost-effective, potential anti-HIV drug with easy availability from the other sources rather than the synthetic source. The natural compounds may prove as the promising alternative options to develop a new anti-HIV drug with negligible side effects and higher efficacy. Various phytochemicals such as curcumin, geranin, gallotannin, tiliroside, kaempferol-3-o-glucoside and trachelogenin etc. have been evaluated for the in-silico analysis against HIV-1 21. The aim of molecular docking is to estimate the realistic binding geometries of a proposed ligand with a target with known target site. Characterization of the binding site, appropriate orientation of the ligand into the binding site and estimation of the strength of interaction of ligand-receptor complex is the main task of the molecular docking. Recently most of the drugs in the market are either derived from the natural sources or semi-synthetic in nature 21. Some analogues of the natural compounds for some new targets in HIV-1 may be developed as the new anti HIV drugs. Since metals and metal-complexes have been used to cure the several critical diseases, therefore, metal-based therapy may be the next step to treat the HIV/AIDS 22. The use of phytochemicals to be explored as anti-HIV agents could be another viable option.
Carissa carandas Linn. (family; Apocynaceae) is commonly known as Karonda. C. carandas, commonly present in India and other Asian countries 23. Fruits and leaves are known for presence of various pharmacologically important phytochemicals 24. The extract of different parts of this plant is used for the cure of various ailments. From the extracts of different parts of C. carandas few potential phytocompounds named as carandinol, carrinsone, carindone, lupeol etc. were identified and isolated 25, 26. In the present paper, we have presented the qualitative characterization of different extracts of leaves and fruits of C. carandas using solvents of varying polarities such as hexane, ethanolic, methanolic, and aqueous. Using some known molecules from this plant as ligands and HIV-1 RT and HIV-1 protease as targets, the in-silico analysis has been carried out in order to assess their prospective applications as effective anti-HIV-1 regimen in future.
Materials and Methods
Collection of Plant Materials
Mature fruits and healthy leaves of C. carandas were collected during November from the Pratapgarh district of Uttar Pradesh, India. Plant materials were washed under running tap water followed by double distilled water and dried under shade condition for 2 h in the hot air oven at 45°C before grinding. The dried material was used to make powder using electrical grinder.
Preparation of Plant Extracts and Qualitative Characterization of Phytochemicals
Compounds Used for Molecular Docking Study Against HIV-1 RT and HIV-1 Proteases
Total nine phytocompounds of C. carandas were used for docking study. The details of the used compounds are mentioned in the Table 1.
Table 1. Compounds used for docking study| Compound name | Common name | Pubchem CID no. | Molecular formula | Chemical name (IUPAC name) |
| Carindone | Not any | 101316738 | C 31 H 44 O 6 | 7,7'-bis(2-hydroxypropan-2-yl)-1',4'a,5,9a-tetramethylspiro3a6, 7, 8, 99b benzofuran-2,3'-5,6,7,8-tetrahydro-4H-naphthalene]-2',3,4-trione |
| Carrisone | Carrison | 5086419 | C15H27O2 | 7-(2-hydroxypropan-2-yl)-1,4a-dimethyl-3,4,5,6,7,8 hexahydronaphthalen-2-one |
| Carandinol | Not any | 102202376 | C 30 H 52 O 2 | (3R,3aR,5aR,5bR,7aR,9S,11aR,11bR,13aR,13bR)-5a,5b,8,8,11a,13b-hexamethyl-3-propan-2-yl 2,3a,4,5,6,7,7a,9,10,11,11b,12,13,13a-tetradecahydro-1H cyclopentaachrysene-3,9-diol |
| Lupeol | Fagarasterol, Clerodol, Monogynol B, | 259846 | C30H50O | 1R,3aR,5aR,5bR,7aR,9S,11aR,11bR,13aR,13bR)-3a,5a,5b,8,8,11a-hexamethyl-1-prop-1-en-2-yl-1,2,3,4,5,6,7,7a,9,10,11,11b,12,13,13a,13bhexadecahydrocyclopentaachrysen-9-ol |
| Scopoletin | Gelseminic acid | 5280460 | C10H8O4 | 7-hydroxy-6-methoxychromen-2-one |
| Ursolic acid | Malol, PrunolUrson, | 64945 | C30H48O3 | (1S,2R,4aS,6aR,6aS,6bR,8aR,10S,12aR,14bS)-10-hydroxy-1,2,6a,6b,9,9,12a-heptamethyl 2,3,4,5,6,6a,7,8,8a,10,11,12, 13, 14b-tetradecahydro-1H-picene-4a-carboxylic acid |
| p-caumaric acid | 637542 | C 9 H 8 O 3 | (E)-3-(4-hydroxyphenyl)prop-2-enoic acid | |
| Rutin | Quercetin 3 Rutinoside | 5280805 | C 27 H 30 O 16 | 2-(3,4-dihydroxyphenyl)-5,7-dihydroxy-3-(2S3R4S5S6R)oxymethyl]oxan-2 yl]oxychromen-4-one |
| Gallic acid | Gallate; Benzoic acid, | 370 | C 7 H 6 O 5 | 3,4,5-Trihydroxybenzoic acid |
Average Linkage Between the Compounds
We used hierarchical clustering facility of ChemMine (webserver) ChemMine tools: an online service for analyzing and clustering small molecules (Nucleic Acids Research, 39 (2011) W486-W491) for clustering of all the phyto-compounds. Atom pair structural descriptors and average linkage method were used for the agglomerative hierarchical clustering with tanimoto coefficient as the similarity measure.
Molecular Docking Methods of Phytochemicals with HIV-1 Molecule
The three dimensional structures of HIV reverse transcriptase (PDB ID: 1REV) and HIV protease (PDB ID: 1EBY) were retrieved from protein data bank at resolutions of 2.6 Å and 2.29 Å, respectively 30. The ligand molecules were retrieved from PubChem database 31. Docking studies were performed by AutoDock 32. For docking experiments with AutoDock, the ligands were preprocessed by addition of Gasteiger partial charges and by merging of non-polar hydrogen atoms with the help of AutoDock tools. Autogrid program was used to create the Affinity maps with 0.375 spacing. Each docking experiment was obtained from ten different runs and simulations were performed using the Lamarckian genetic algorithm (LGA). The docking experiments by AutoDock were both focused on the docking site on the predefined binding site by ligand TB9 (9-chloro-TIBO) in 1REV and by BAE369 in 1EBY.
Results
Phytochemical Screening
Qualitative characterization of the leaf and fruit extracts showed the presence of various phytocompounds. C. carnadas extracts showed the presence of carbohydrates, fat, oil, phenols, flavonoids, alkaloids, terpenoids, steroids, glycosides, saponins, glycosides, and tannins (Table 2). Some compounds were present in very little amount and some were in excess. Ethanolic and methanolic extracts showed the presence of most of the phytochemicals while hexane and aqueous extracts displayed lesser amount of phytochemicals.
Table 2. Efficiency ranking and molecular docking data of phytochemicals against 1REV| Ligands | Binding energy(Kcal/mole) | KI | Intermolecular energy(Kcal/mole) | Internal energy(Kcal/mole) | Torsional energy(Kcal/mole) | Efficiency ranking |
| Carandinol | -10.11 | 39.14nM | -11.00 | -0.16 | 0.89 | 1 |
| Carindone | -8.85 | 324.25nM | -10.05 | -0.52 | 1.19 | 2 |
| Carinsone | -8.71 | 409.19nM | -9.31 | 0.25 | 0.6 | 3 |
| Lupeol | -7.54 | 2.97uM | -8.14 | -0.16 | 0.6 | 4 |
| Ursolic acid | -7.47 | 3.37uM | -8.36 | 0.33 | 0.89 | 5 |
| Gallic acid | -6.58 | 14.94uM | -11.36 | -8.78 | 4.77 | 6 |
| p-coumaric acid | -4.64 | 395.5uM | -5.84 | -0.08 | 1.19 | 7 |
| Scopoletin | -4.88 | 265.12uM | -5.48 | -0.23 | 0.6 | 8 |
| Rutin | -4.41 | 585.02uM | -9.18 | -4.67 | 4.77 | 9 |
Average Linkage Between the Compounds
The average linkages have been shown in the hierarchal study of all nine compounds Figure 1. Rutin and carrinsone are more structurally similar. Carandinol and lupeol are are similar but derived from the ursolic acid. Scopoletin and p-coumaric acid are similar to each other and structurally derived from the gallic acid.
Figure 1.Hierarchical clustering of all the phyto-compounds present (Average linkage method)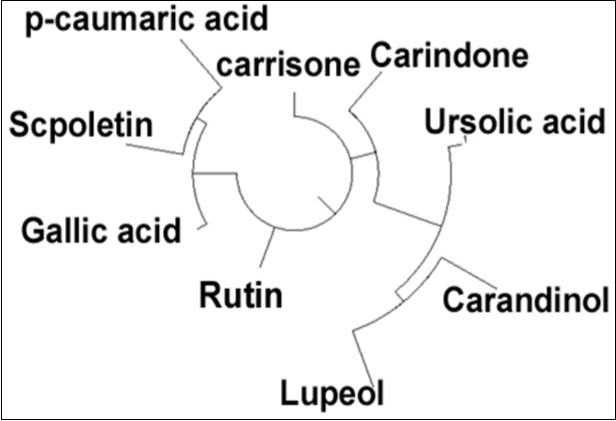
Docking Study of the Compounds Against 1REV and 1EBY
Molecular docking was performed to analyze the inhibitory action of each phytocompound with both HIV-1 reverse transcriptase (1REV) and HIV-1 protease (1EBY). To find the best binding pose of ligand at the active site of protein, the binding energy was calculated by adding the final intermolecular energy and torsional free energy. Our study revealed that Carandinol, Carindone and Carinsone docked against HIV-1 reverse transcriptase and HIV-1 protease showed maximum binding with binding energy; the values being 10.11, -8.85, -8.71 kcal/mole and -10.99, -9.38 and -7.23 kcal/mole, respectively.
Carnadinol was most effective compound against both 1REV (Table 3) and 1EBY. Rutin was least effective against 1REV while p-coumaric acid was least effective against 1EBY. 1REV contains several active sites. The active site of 1REV comprises leucine, glycine, tryptophan, tyrosine valine, phenylalanine, and alanine etc. Carandinol dominantly binds with LEU100, Val106, Val179, ILE180, TYR181, TYR188, PHE 227, LEU234, TYR318 and inhibits the activity of 1REV. While p-coumaric acid has least binding affinity toward the active sites and it only binds with few amino acids like GLN23, LYS 22, PRO59, ASP76, PHE61, TRP24, GLU399 and showed least inhibition. Molecular docking between the ligands and target molecule has been clearly demonstrated in Figure 2.
Table 3. Efficiency ranking and molecular docking data of phytochemicals against 1EBY| Ligands | Binding energy(Kcal/mole) | KI | Intermolecular energy(Kcal/mole) | Internal energy(Kcal/mole) | Torsional energy(Kcal/mole) | Efficiency ranking |
| Carandinol | -10.99 | 8.87nM | -11.88 | 0.21 | 0.89 | 1 |
| Carindone | -9.38 | 133.79nM | -10.57 | 0.52 | 1.19 | 3 |
| Carinsone | -7.23 | 5.0uM | -7.83 | -0.26 | 0.6 | 6 |
| Lupeol | -10.24 | 31.14nM | -10.84 | -0.26 | 0.6 | 2 |
| Ursolic acid | -8.37 | 738.3nM | -9.26 | 0.07 | 0.89 | 4 |
| Gallic acid | -8.01 | 1.35uM | -12.78 | -5.2 | 4.77 | 5 |
| p-coumaric acid | -4.7 | 357.15uM | -5.9 | -0.08 | 1.19 | 9 |
| Scopoletin | -5.67 | 69.4uM | -6.27 | -0.13 | 0.6 | 8 |
| Rutin | -6.28 | 24.76uM | -11.06 | -4.84 | 4.77 | 7 |
Figure 2.Protein-ligand interactions: (a) Indicates interaction between HIV-1 RT and carandinol (b) Indicates interaction between HIV-1 protease and carandinol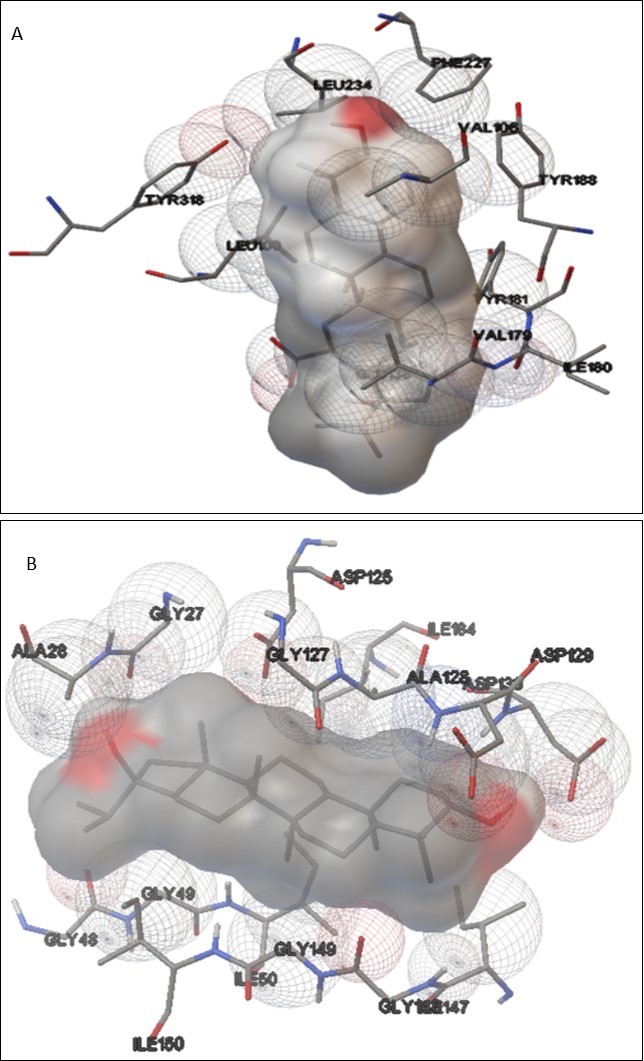
Discussion
The plant extracts of C. carandas showed the presence of many phytochemicals. Phytochemical screening of the leaves and root extracts of this plant has been reported by Sawant and Godghate 19. Qualitative characterization has also been reported in other plants and presence of phytochemicals has been also reported in other plants 29, 33.
Carandinol showed the highest inhibition activity and rutin the least inhibition activity against both 1REV and 1 EBY. The inhibition potential of ligand depends upon the active sites of molecules and structure of ligands. Molecular docking was also performed by other worker against HIV-1 RT 21 and HIV-1 integrase 34 and reported the significant anti-HIV activity of phytochemicals. In-silico analysis of Dolichin A and B against HIV-RT was also evaluated 35.
Conclusions
From the present study several compounds were screened in the leaf and fruit extracts. In this work, molecular docking was performed by phytocompounds with both HIV-1 reverse transcriptase and HIV-1 protease. Based on the 3D structure obtained from PDB, the molecular docking was used to produce the binding pose of best docked structure. Total nine compounds were used for molecular docking and among all the carandinol exhibited to be the most potential inhibitor against 1REV and 1EBY. Based on the molecular docking results, it was evident that carandinol possessed lowest binding energy of -10.11 kcal/mole with HIV-1 reverse transcriptase and -10.99 kcal/mole with HIV-1 protease. Inhibition of 1REV and 1EBY proved that the plant extract of C. carandas had the anti-HIV potential. Hence, it can be exploited to develop more effective and potential anti-HIV drugs.
Contribution
RS and BS conceived the idea on subject of research mentioned in the manuscript. RS performed the phytochemical characterization. RS and AN performed the docking study. All the authors have contributed in preparation and editing of the manuscript.
Acknowledgements
RS is grateful to DST-New Delhi for financial support in the form of N-PDF (PDF/2016/000061). BS is grateful to UPCST-Lucknow for financial support in the form of a research grant.
References
- 2.Douek D C, Roederer M.. Koup RA.(2009)Emerging Concepts in the Immunopathogenesis of AIDS, Annu Rev Med.60 471.
- 3.Sharma B. (2014) Oxidative stress in HIV Pateints receiving Antiretroviral therapy. , Current HIV-Research 12, 13-21.
- 5.HemelaarJ. (2012) The origin and diversity of the HIV-1 pandemic. , Trends in Molecular Medicine 18, 182-192.
- 9.Boyer P L, Coffin J M, Delviks-Frankenberry K A. (2008) HIV-1 reverse transcriptase connection subdomain mutations reduce template RNA degradation and enhance AZT excision.Proc. , Natl. Acad. Sci. U.S.A 105, 10943-10948.
- 10.Giridhar R, Prajapati D G, Ramajayam R, Yadav M R.(2009)The search for potent, small molecule NNRTIs: A review. Bioorganic Medicinal Chemistry.17 5744-5762.
- 11.Anderson K S.(2007).Reverse transcription of the HIV-1 pandemic. , The FASEB Journal 21, 3795-3808.
- 12.Herschorn A, Hizi. (2008) Retroviral reverse transcriptases (other than those of HIV-1 and murine leukemia virus): A comparison of their molecular and biochemical properties.Virus Res. 134, 203-20.
- 13.Gu W-G, Zhang X.. Yuan JF.(2014) Anti-HIV Drug Development through Computational Methods.The AAPSJ 16, 674-80.
- 14.Huang X, Britto M D, Kear-Scott J L, Boone C D, Rocca J R.et al.(2014) The role of select subtype polymorphisms on HIV-1 protease conformational sampling and dynamics.Journal ofBiological. Chemistry,289 17203-17214.
- 15.Kohl N E, Emini E A, Schleif W A. (1988) Active human immunodeficiency virus protease is required for viral infectivity. , Proceeding of National Academy of Science 85, 4686-90.
- 16.Krausslich H G, Ingraham R H, Skoog M T, Wimmer E, Pallai P V. (1989) Activity of purified biosynthetic proteinase of human immunodeficiency virus on natural substrates and synthetic peptides". , Proceeding of National Academy of Science U.S.A 86, 807-11.
- 17.Seelmeier S, Schmidt H, Turk V, Helm K von der. (1988) Human immunodeficiency virus has an aspartic-type protease that can be inhibited by pepstatin A. Proceeding of. , National Academy of Science U.S.A 85, 6612-6.
- 18.McPhee F, Good A C, Kuntz I D, Craik C S. (1996) Engineering human immunodeficiency virus 1 protease heterodimers as macromolecular inhibitors of viral maturation. , Proc. Natl. Acad. Sci. U.S.A 93, 11477-81.
- 19.Clercq E D. (2009) Anti-HIV drugs: 25 compounds approved within 25 years after the discovery of HIV. , International Journal of Antimicrobial Agent 33, 307-320.
- 21.Seal A, Aykkal R, Babu R O, Ghosh M. (2011) Docking study of HIV-1 reverse transcriptase with phytochemicals. , Bioinformation 5, 430-9.
- 22.Singh R, Sharma B. (2018) . Metal-Based Therapy in Traditional and Modern Medicine Systems inBiomedical Applications of Metals 195-211.
- 23.Sawant R S, Godghate A G. (2013) Comparative studies of phytochemical screening ofCarissacarandasLinn. , Asian Journal of Plant Science Research.3 21-25.
- 24.Panda D, Panda S, Pramanik K, MondalS. (2014) Karonda (Carissaspp.): An Underutilized Minor Fruit Crop with Therapeutic and Medicinal Use. , International Journal of Economic Plants 1, 009-014.
- 25.Begum S, Syed S A, Siddiqui B S, Sattar S A, ChoudharyMI.(2013)Carandinol: First isohopanetriterpene from the leaves ofCarissa carandasL. and its cytotoxicity against cancer cell lines. Phytochemistry. Letters.6 91-95.
- 26.Ganapaty S, Laatsch H BharathCH. (2010) Des-N-Methylnoracronycine from the roots ofCarissaconjesta.Wight. , International Journal of Green Pharmacy 186-188.
- 27.Singh R, Kumari N. (2015) Comparative determination of phytochemicals and antioxidant activity from leaf and fruit ofSapindusmukorossiGaertn. – A valuable medicinal tree, Industrial Crops and. Products.73 1-8.
- 28.Harborne J B. (1973) A guide to Modern techniques of plant analysis; phytochemical methods, London, Chapman and Hall Ltd. 49-188.
- 29.Singh R, Kumari N, Gangwar M, Nath G. (2015) Qualitative characterization of phytochemicals andin vitroantimicrobial evaluation of leaf extract ofCouroupitaguianensisAubl. – a threatened medicinal tree. , International Journal of Pharmacy and Pharmaceutical Sciences 7, 212-215.
- 30.Berman H M, Westbrook J, Feng Z. (2000) The Protein Data Bank. , Nucleic Acids Research 28, 235-242.
- 31.Wang Y, Xiao J, Suzek T O, Zhang J, Wang J et al. (2009) PubChem: a public information system for analyzing bioactivities of small molecules. , Nucleic Acids Research 37, 623-633.
- 32.Morris G M, Huey R, Lindstrom W. (2009) AutoDock4 and AutoDockTools4: Automated Docking with Selective Receptor Flexibility. , Journal of Computational Chemistry.30 2785-2791.
- 33.Singh R, Kumari N.Nath G.(2016)Free radical scavenging activity and antimicrobial potential of leaf and fruit extractsofSapindusmukorossiGaertn. against clinical pathogens. , International Journal of Phytomedicine 8, 22-28.
Cited by (4)
This article has been cited by 4 scholarly works according to:
Citing Articles:
Bioinformatics and Biology Insights (2022) OpenAlex
Bioinformatics and Biology Insights (2022) Crossref
E. M. Terefe, Arabinda Ghosh - Bioinformatics and Biology Insights (2022) Semantic Scholar
Springer eBooks (2021) OpenAlex
O. Oluwole, O. Ademuyiwa - Antioxidants in Plant-Microbe Interaction (2021) Semantic Scholar
